Compound details
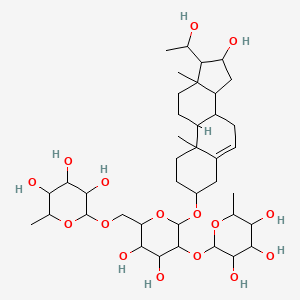
Balagyptin
| Compound ID | CDAMM03018 |
|---|---|
| Common name | Balagyptin | IUPAC name | 2-[[3,4-dihydroxy-6-[[16-hydroxy-17-(1-hydroxyethyl)-10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-3-yl]oxy]-5-(3,4,5-trihydroxy-6-methyloxan-2-yl)oxyoxan-2-yl]methoxy]-6-methyloxane-3,4,5-triol |
| Molecular formula | C39H64O16 |
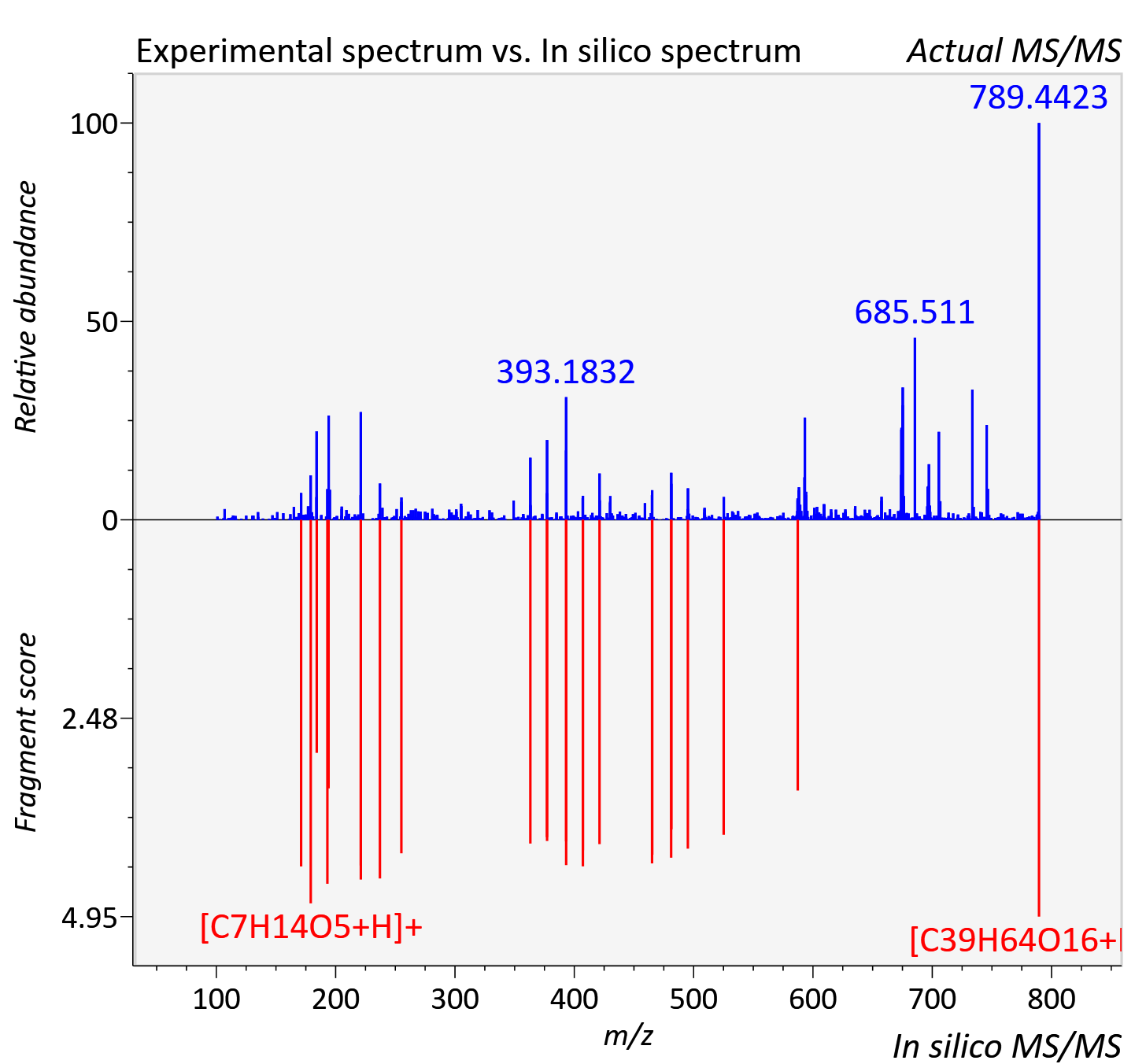
Experimental data
| Retention time | 17.6 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 789.436 | Theoretical mz | 789.426 |
| Error | 12.7 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.7846 |
Identifiers and class information
| Inchi key | DHEBGTQGALZORI-UHFFFAOYNA-N |
|---|---|
| Smiles | OC(C)C1C(O)CC2C3CC=C4CC(OC5OC(COC6OC(C)C(O)C(O)C6O)C(O)C(O)C5OC7OC(C)C(O)C(O)C7O)CCC4(C)C3CCC21C |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q15125 | EBP | Anti-estrogen binding site (AEBS) | T25317 | SEA |
| P31213 | SRD5A2 | Steroid 5-alpha-reductase 2 | T71390 | SEA |
| P08185 | SERPINA6 | Corticosteroid binding globulin | T88452 | SEA |
| Q9UHC9 | NPC1L1 | Niemann-Pick C1-like protein 1 | T33901 | SEA |
| Q13133 | NR1H3 | LXR-alpha | T52297 | SEA |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
| P05093 | CYP17A1 | Cytochrome P450 17A1 | T89041 | SEA |
| P05230 | FGF1 | Acidic fibroblast growth factor | T18639 | SEA |
| P60568 | IL2 | Interleukin-2 | T61698 | SwissTargetPrediction and SEA |
| P04746 | AMY2A | Pancreatic alpha-amylase | T86918 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
| O15439 | ABCC4 | Multidrug resistance protein 4 | T39919 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T25317 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q15125 | EBP |
| T71390 | DI0348 | Prostate hyperplasia | [ICD-11: GA90] | P31213 | SRD5A2 |
| T88452 | DI0037 | Asthma | [ICD-11: CA23] | P08185 | SERPINA6 |
| T88452 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P08185 | SERPINA6 |
| T88452 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08185 | SERPINA6 |
| T33901 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | Q9UHC9 | NPC1L1 |
| T52297 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | Q13133 | NR1H3 |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T89041 | DI0346 | Prostate cancer | [ICD-11: 2C82] | P05093 | CYP17A1 |
| T18639 | DI0081 | Chronic arterial occlusive disease | [ICD-11: BD4Z] | P05230 | FGF1 |
| T18639 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P05230 | FGF1 |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T86918 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P04746 | AMY2A |
| T86918 | DI0328 | Pancreatic malfunction | [ICD-11: DC30-DC3Z] | P04746 | AMY2A |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
