Compound details
Lanceolitol A7
| Compound ID | CDAMM03011 |
|---|---|
| Common name | Lanceolitol A7 | IUPAC name | [2,3,4,5-tetrahydroxy-6-(3,4,5-trihydroxyoxan-2-yl)oxycyclohexyl] icosanoate |
| Molecular formula | C31H58O11 |
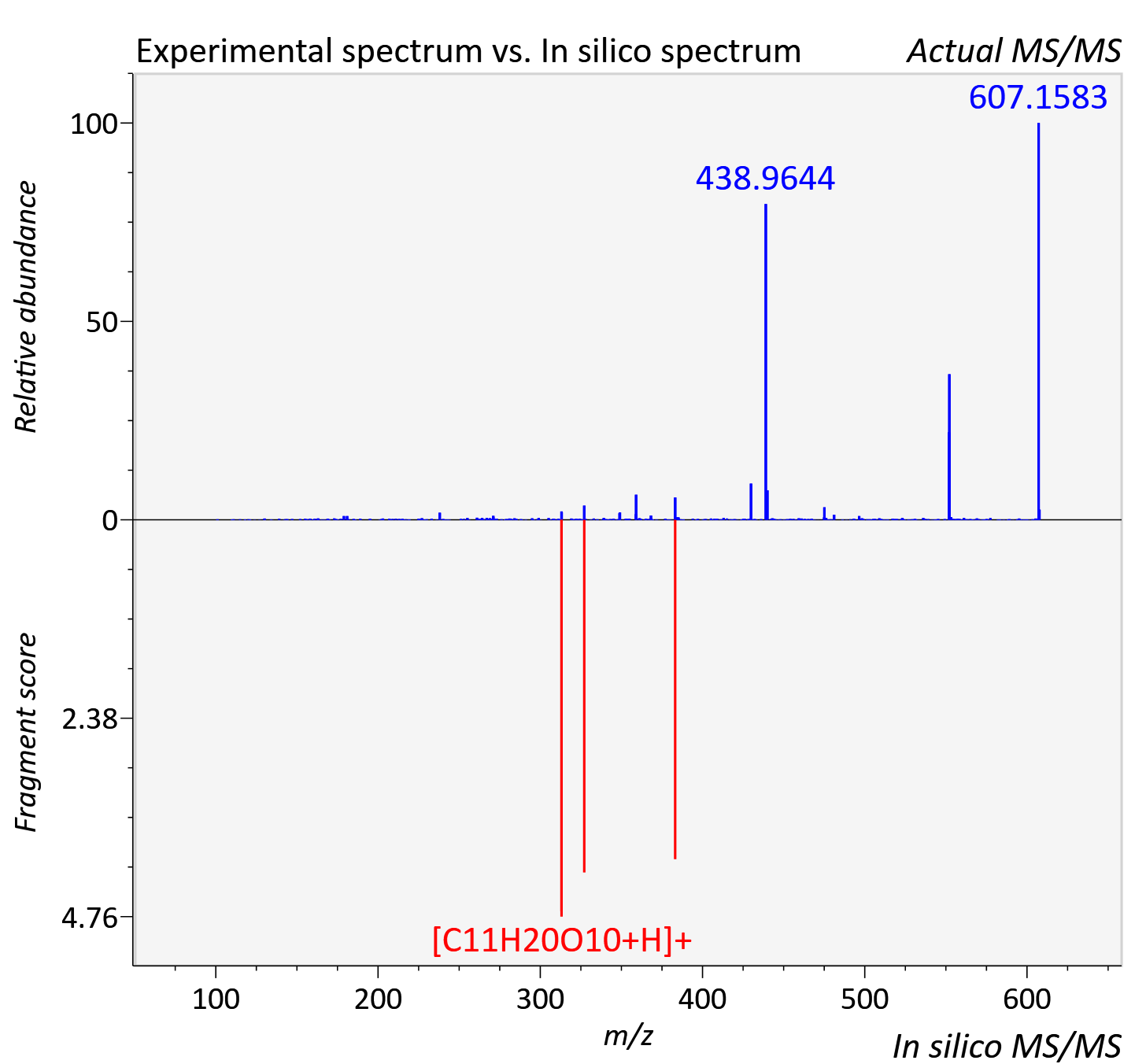
Experimental data
| Retention time | 19.45 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 607.41 | Theoretical mz | 607.405 |
| Error | 7.61 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.9527 |
Identifiers and class information
| Inchi key | XBMWPUJNCJDGGT-YJOSPMTENA-N |
|---|---|
| Smiles | O=C(OC1C(O)C(O)C(O)C(O)C1OC2OCC(O)C(O)C2O)CCCCCCCCCCCCCCCCCCC |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SEA |
| P21554 | CNR1 | Cannabinoid receptor 1 (by homology) | T76685 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P25021 | HRH2 | Histamine H2 receptor | T30985 | SEA |
| P35367 | HRH1 | Histamine H1 receptor | T77913 | SEA |
| O00206 | TLR4 | Toll-like receptor 4 (by homology) | T81443 | SEA |
| P10827 | THRA | Thyroid hormone receptor alpha | T79591 | SEA |
| P10828 | THRB | Thyroid hormone receptor beta-1 | T98933 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P05230 | FGF1 | Acidic fibroblast growth factor | T18639 | SEA |
| P09038 | FGF2 | Basic fibroblast growth factor | T31621 | SEA |
| P34972 | CNR2 | Cannabinoid receptor 2 | T37693 | SEA |
| P16109 | SELP | P-selectin | T10965 | SEA |
| Q9H228 | S1PR5 | Sphingosine 1-phosphate receptor Edg-8 | T50089 | SEA |
| P40189 | IL6ST | Interleukin-6 receptor subunit beta | T93440 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| O00142 | TK2 | Thymidine kinase, mitochondrial | T22976 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
| Q9UMJ8 | GBA1 | Lysosomal acid glucosylceramidase | T63243 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T76685 | DI0031 | Anorexia nervosa | [ICD-11: 6B80] | P21554 | CNR1 |
| T76685 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P21554 | CNR1 |
| T76685 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P21554 | CNR1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T30985 | DI0161 | Gastro-oesophageal reflux disease | [ICD-11: DA22] | P25021 | HRH2 |
| T30985 | DI0313 | Oesophageal/gastroduodenal disorder | [ICD-11: DD90] | P25021 | HRH2 |
| T30985 | DI0333 | Peptic ulcer | [ICD-11: DA61] | P25021 | HRH2 |
| T77913 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | P35367 | HRH1 |
| T77913 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | P35367 | HRH1 |
| T77913 | DI0063 | Breathing abnormality | [ICD-11: MD11] | P35367 | HRH1 |
| T77913 | DI0099 | Conjunctiva disorder | [ICD-11: 9A60] | P35367 | HRH1 |
| T77913 | DI0105 | Cough | [ICD-11: MD12] | P35367 | HRH1 |
| T77913 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P35367 | HRH1 |
| T77913 | DI0136 | Episodic vestibular syndrome | [ICD-11: AB31] | P35367 | HRH1 |
| T77913 | DI0173 | Headache | [ICD-11: 8A80-8A84] | P35367 | HRH1 |
| T77913 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P35367 | HRH1 |
| T77913 | DI0272 | Morning sickness disorder | [ICD-11: SC00] | P35367 | HRH1 |
| T77913 | DI0292 | Nasopharyngitis | [ICD-11: CA00] | P35367 | HRH1 |
| T77913 | DI0293 | Nausea/vomiting | [ICD-11: MD90] | P35367 | HRH1 |
| T77913 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P35367 | HRH1 |
| T77913 | DI0349 | Pruritus | [ICD-11: EC90] | P35367 | HRH1 |
| T77913 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P35367 | HRH1 |
| T77913 | DI0388 | Sleep-wake disorder | [ICD-11: 7A00-7B2Z] | P35367 | HRH1 |
| T77913 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | P35367 | HRH1 |
| T81443 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | O00206 | TLR4 |
| T81443 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O00206 | TLR4 |
| T81443 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | O00206 | TLR4 |
| T79591 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10827 | THRA |
| T79591 | DI0197 | Hypo-thyroidism | [ICD-11: 5A00] | P10827 | THRA |
| T98933 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P10828 | THRB |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T18639 | DI0081 | Chronic arterial occlusive disease | [ICD-11: BD4Z] | P05230 | FGF1 |
| T18639 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P05230 | FGF1 |
| T31621 | DI0005 | Acne vulgaris | [ICD-11: ED80] | P09038 | FGF2 |
| T37693 | DI0040 | Attention deficit hyperactivity disorder | [ICD-11: 6A05] | P34972 | CNR2 |
| T37693 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P34972 | CNR2 |
| T10965 | DI0088 | Circulatory system disease | [ICD-11: BE2Z] | P16109 | SELP |
| T10965 | DI0381 | Sickle-cell disorder | [ICD-11: 3A51] | P16109 | SELP |
| T50089 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | Q9H228 | S1PR5 |
| T93440 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P40189 | IL6ST |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
| T22976 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | O00142 | TK2 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
| T63243 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | Q9UMJ8 | GBA1 |
