Compound details
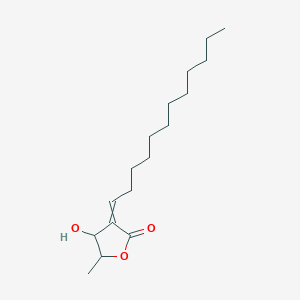
3-Epilitsenolide D1
| Compound ID | CDAMM02999 |
|---|---|
| Common name | 3-Epilitsenolide D1 | IUPAC name | 3-dodecylidene-4-hydroxy-5-methyloxolan-2-one |
| Molecular formula | C17H30O3 |
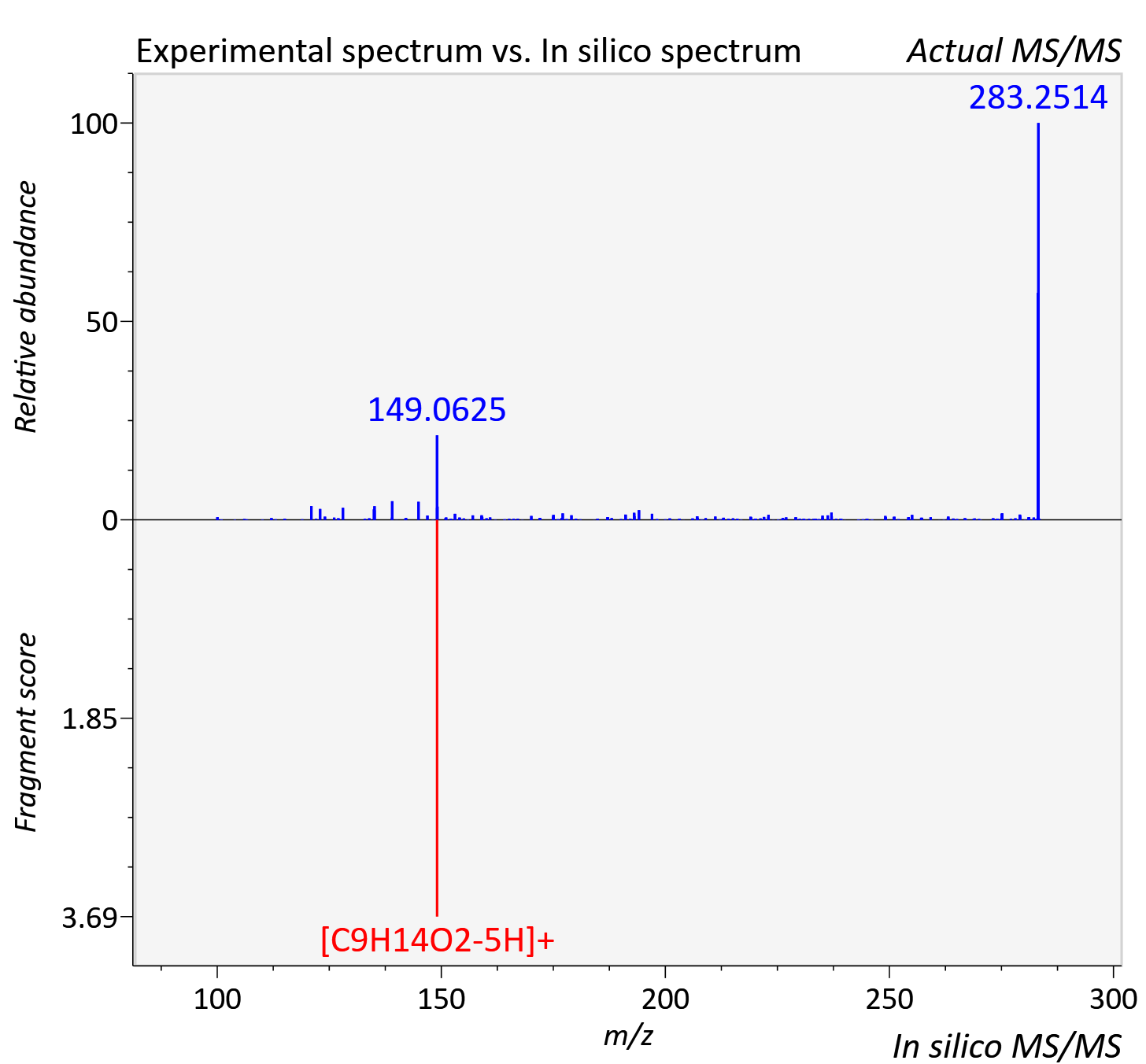
Experimental data
| Retention time | 14.91 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 283.223 | Theoretical mz | 283.226 |
| Error | 12.63 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.6832 |
Identifiers and class information
| Inchi key | NSMMPEJLOMMKCX-VYCBQPGPNA-N |
|---|---|
| Smiles | O=C1OC(C)C(O)C1=CCCCCCCCCCCC |
| Superclass | Organoheterocyclic compounds |
| Class | Lactones |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| Q02156 | PRKCE | Protein kinase C epsilon | T00895 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T00895 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | Q02156 | PRKCE |
| T00895 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q02156 | PRKCE |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
