Compound details
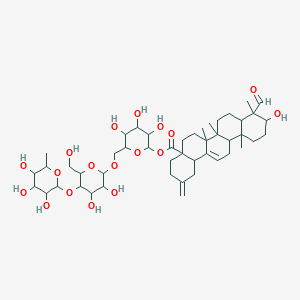
Acanjaposide B
| Compound ID | CDAMM02970 |
|---|---|
| Common name | Acanjaposide B | IUPAC name | [6-[[3,4-dihydroxy-6-(hydroxymethyl)-5-(3,4,5-trihydroxy-6-methyloxan-2-yl)oxyoxan-2-yl]oxymethyl]-3,4,5-trihydroxyoxan-2-yl] 9-formyl-10-hydroxy-6a,6b,9,12a-tetramethyl-2-methylidene-1,3,4,5,6,6a,7,8,8a,10,11,12,13,14b-tetradecahydropicene-4a-carboxylate |
| Molecular formula | C47H72O18 |
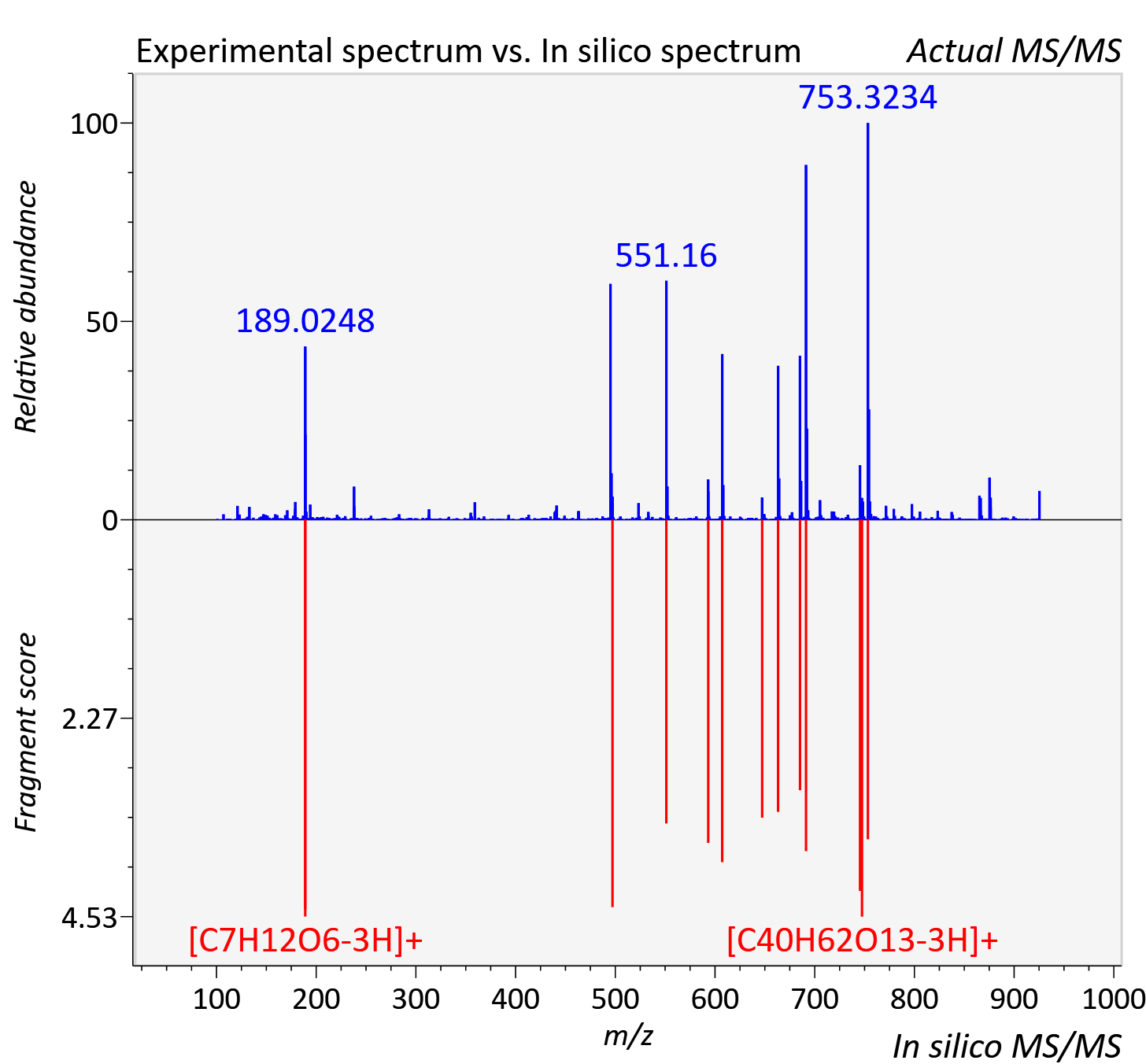
Experimental data
| Retention time | 21.84 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 925.488 | Theoretical mz | 925.479 |
| Error | 9.32 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.3764 |
Identifiers and class information
| Inchi key | MLWNDZPJHCBVPM-RWKSZYQRNA-N |
|---|---|
| Smiles | O=CC1(C)C(O)CCC2(C)C3CC=C4C5CC(=C)CCC5(C(=O)OC6OC(COC7OC(CO)C(OC8OC(C)C(O)C(O)C8O)C(O)C7O)C(O)C(O)C6O)CCC4(C)C3(C)CCC12 |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P29350 | PTPN6 | Protein-tyrosine phosphatase 1C | T98264 | SEA |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SEA |
| P17706 | PTPN2 | T-cell protein-tyrosine phosphatase | T49156 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| P13726 | F3 | Coagulation factor VII/tissue factor | T72702 | SEA |
| P04746 | AMY2A | Pancreatic alpha-amylase | T86918 | SEA |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T49156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P17706 | PTPN2 |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T72702 | DI0075 | Cervical cancer | [ICD-11: 2C77] | P13726 | F3 |
| T86918 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P04746 | AMY2A |
| T86918 | DI0328 | Pancreatic malfunction | [ICD-11: DC30-DC3Z] | P04746 | AMY2A |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
