Compound details
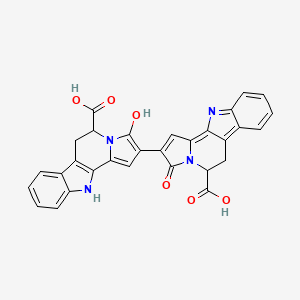
Trichotomine
| Compound ID | CDAMM02936 |
|---|---|
| Common name | Trichotomine | IUPAC name | 2-(5-carboxy-3-hydroxy-6,11-dihydro-5H-indolizino[8,7-b]indol-2-yl)-3-oxo-5,6-dihydroindolizino[8,7-b]indole-5-carboxylic acid |
| Molecular formula | C30H20N4O6 |
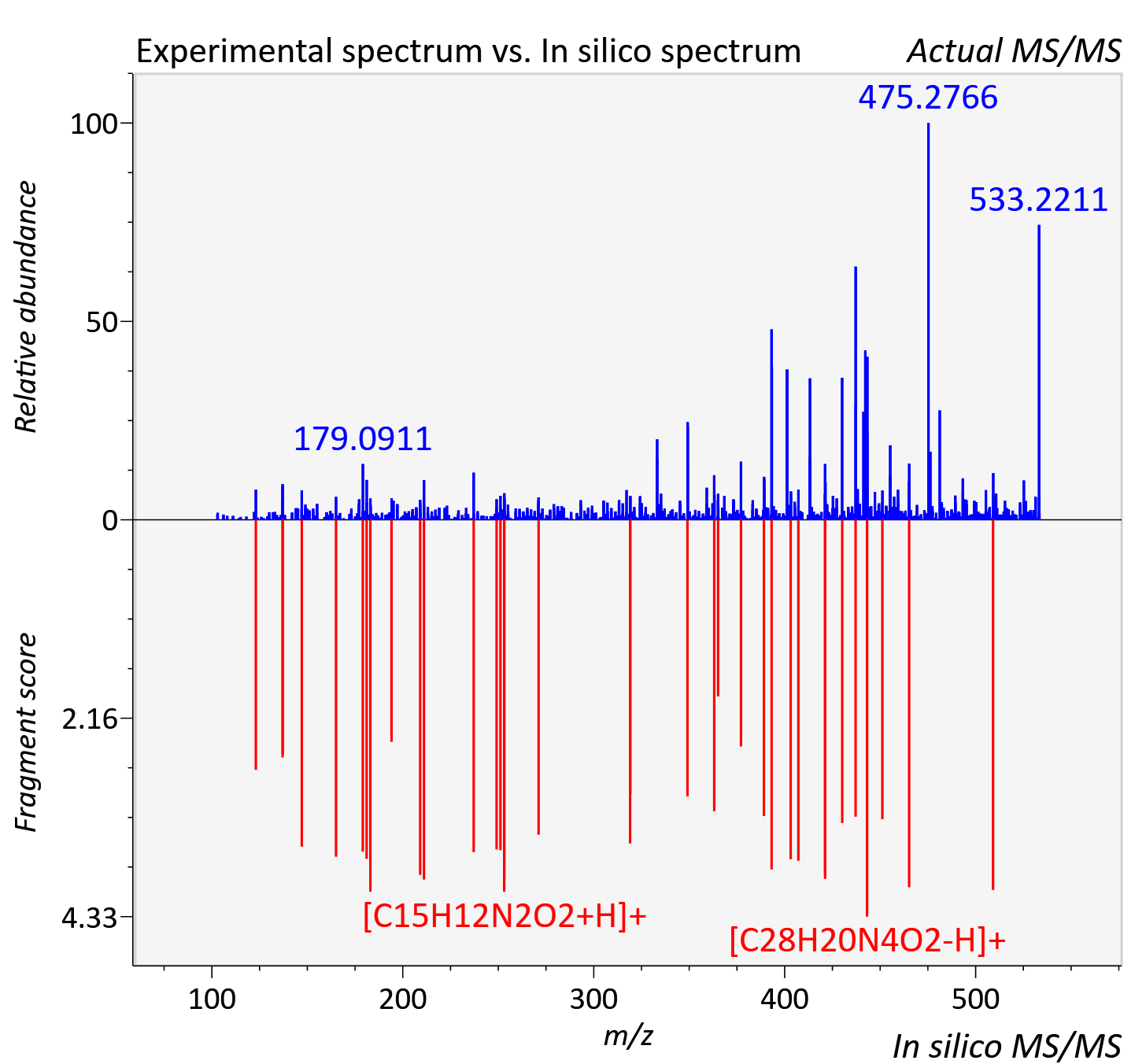
Experimental data
| Retention time | 15.72 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 533.148 | Theoretical mz | 533.145 |
| Error | 4.73 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.9345 |
Identifiers and class information
| Inchi key | LOHPAPKRMNSIDN-PJWPLZPDNA-N |
|---|---|
| Smiles | O=C(O)C1N2C(=O)C(C=C2C=3NC=4C=CC=CC4C3C1)=C5C=C6C=7NC=8C=CC=CC8C7CC(C(=O)O)N6C5=O |
| Superclass | Organoheterocyclic compounds |
| Class | Indoles and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P21918 | DRD5 | Dopamine D5 receptor | T46828 | SEA |
| P52732 | KIF11 | Kinesin-like protein 1 | T28484 | SEA |
| P35346 | SSTR5 | Somatostatin receptor 5 | T64830 | SEA |
| P30874 | SSTR2 | Somatostatin receptor 2 | T53024 | SEA |
| P31391 | SSTR4 | Somatostatin receptor 4 | T62974 | SEA |
| P30872 | SSTR1 | Somatostatin receptor 1 | T16633 | SEA |
| P32745 | SSTR3 | Somatostatin receptor 3 | T13644 | SEA |
| O76074 | PDE5A | Phosphodiesterase 5A | T07663 | SEA |
| P08151 | GLI1 | Zinc finger protein GLI1 | T40890 | SEA |
| Q9HCN6 | GP6 | Platelet glycoprotein VI | T60514 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T46828 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | P21918 | DRD5 |
| T28484 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P52732 | KIF11 |
| T28484 | DI0172 | Head and neck cancer | [ICD-11: 2D42] | P52732 | KIF11 |
| T28484 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P52732 | KIF11 |
| T28484 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P52732 | KIF11 |
| T28484 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | P52732 | KIF11 |
| T28484 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P52732 | KIF11 |
| T28484 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P52732 | KIF11 |
| T28484 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P52732 | KIF11 |
| T64830 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P35346 | SSTR5 |
| T53024 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P30874 | SSTR2 |
| T53024 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P30874 | SSTR2 |
| T53024 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P30874 | SSTR2 |
| T62974 | DI0087 | Chronic pain | [ICD-11: MG30] | P31391 | SSTR4 |
| T62974 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P31391 | SSTR4 |
| T16633 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P30872 | SSTR1 |
| T16633 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P30872 | SSTR1 |
| T13644 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P32745 | SSTR3 |
| T07663 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | O76074 | PDE5A |
| T07663 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | O76074 | PDE5A |
| T07663 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | O76074 | PDE5A |
| T07663 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | O76074 | PDE5A |
| T60514 | DI0035 | Arterial occlusive disease | [ICD-11: BD40] | Q9HCN6 | GP6 |
| T60514 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | Q9HCN6 | GP6 |
