Compound details
Anhydrovariabilin
| Compound ID | CDAMM02927 |
|---|---|
| Common name | Anhydrovariabilin | IUPAC name | 3,9-dimethoxy-6H-[1]benzofuro[3,2-c]chromene |
| Molecular formula | C17H14O4 |
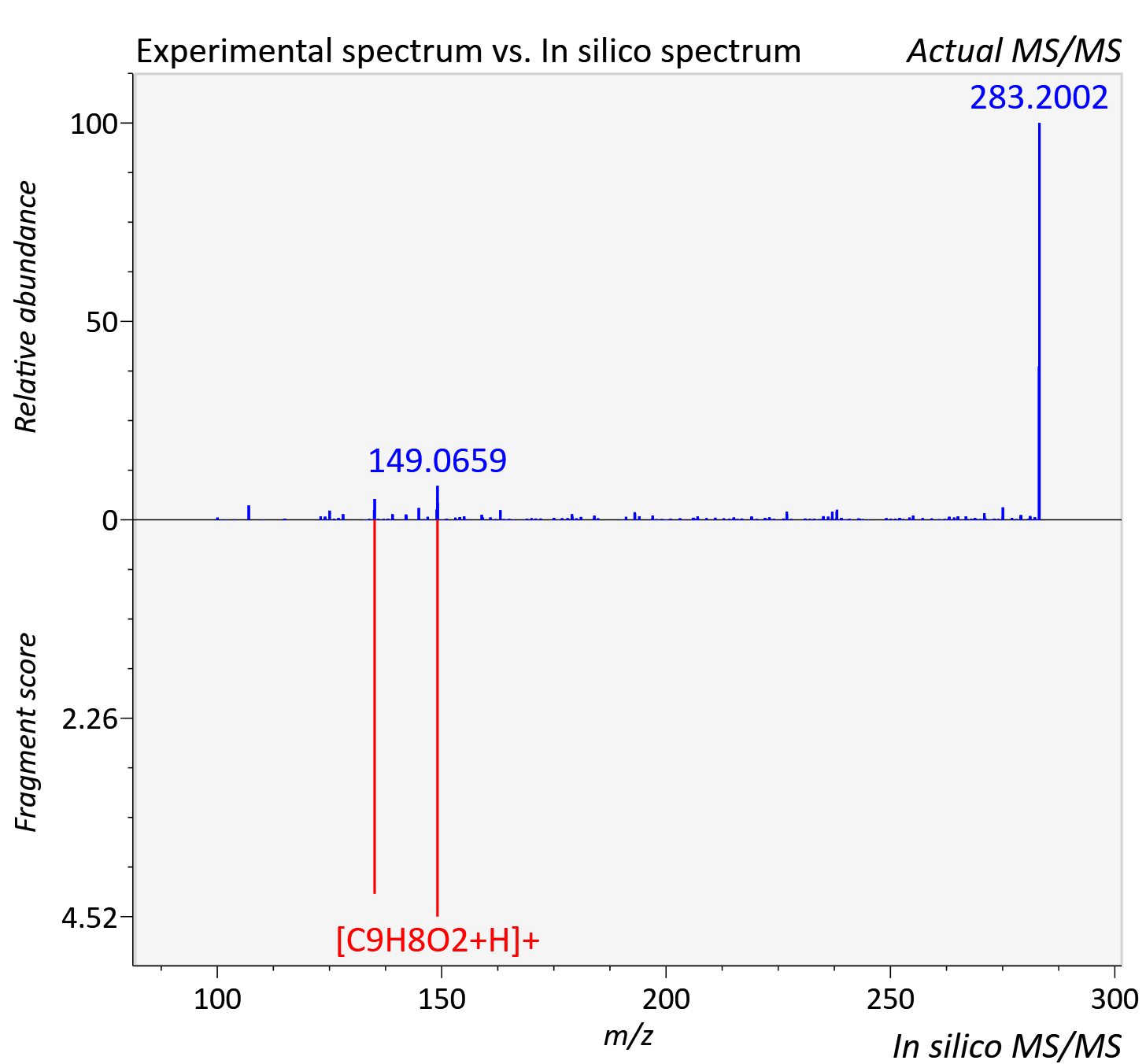
Experimental data
| Retention time | 14.94 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 283.1 | Theoretical mz | 283.096 |
| Error | 14.57 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.8526 |
Identifiers and class information
| Inchi key | FLEXCYTURSFUNC-UHFFFAOYSA-N |
|---|---|
| Smiles | O1C2=CC(OC)=CC=C2C3=C1C=4C=CC(OC)=CC4OC3 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Isoflavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P05067 | APP | Beta amyloid A4 protein | T87024 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SEA |
| P11474 | ESRRA | Estrogen-related receptor alpha | T72841 | SEA |
| O95718 | ESRRB | Estrogen-related receptor beta | T42025 | SEA |
| P08842 | STS | Steryl-sulfatase | T33489 | SEA |
| P48039 | MTNR1A | Melatonin receptor 1A | T97613 | SEA |
| P49286 | MTNR1B | Melatonin receptor 1B | T48268 | SEA |
| Q9NUW8 | TDP1 | Tyrosyl-DNA phosphodiesterase 1 | T33492 | SEA |
| Q9UNQ0 | ABCG2 | ATP-binding cassette sub-family G member 2 | T56556 | SEA |
| P41235 | HNF4A | Hepatocyte nuclear factor 4-alpha | T15514 | SEA |
| Q9H4B7 | TUBB1 | Tubulin beta-1 chain | T84397 | SEA |
| O15427 | SLC16A3 | Monocarboxylate transporter 4 | T31788 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T87024 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P05067 | APP |
| T87024 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P05067 | APP |
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T72841 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P11474 | ESRRA |
| T33489 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | P08842 | STS |
| T97613 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P48039 | MTNR1A |
| T48268 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P49286 | MTNR1B |
| T56556 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9UNQ0 | ABCG2 |
| T56556 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9UNQ0 | ABCG2 |
| T84397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9H4B7 | TUBB1 |
