Compound details
2-Furanmethanol
| Compound ID | CDAMM02908 |
|---|---|
| Common name | 2-Furanmethanol | IUPAC name | furan-2-ylmethanol |
| Molecular formula | C5H6O2 |
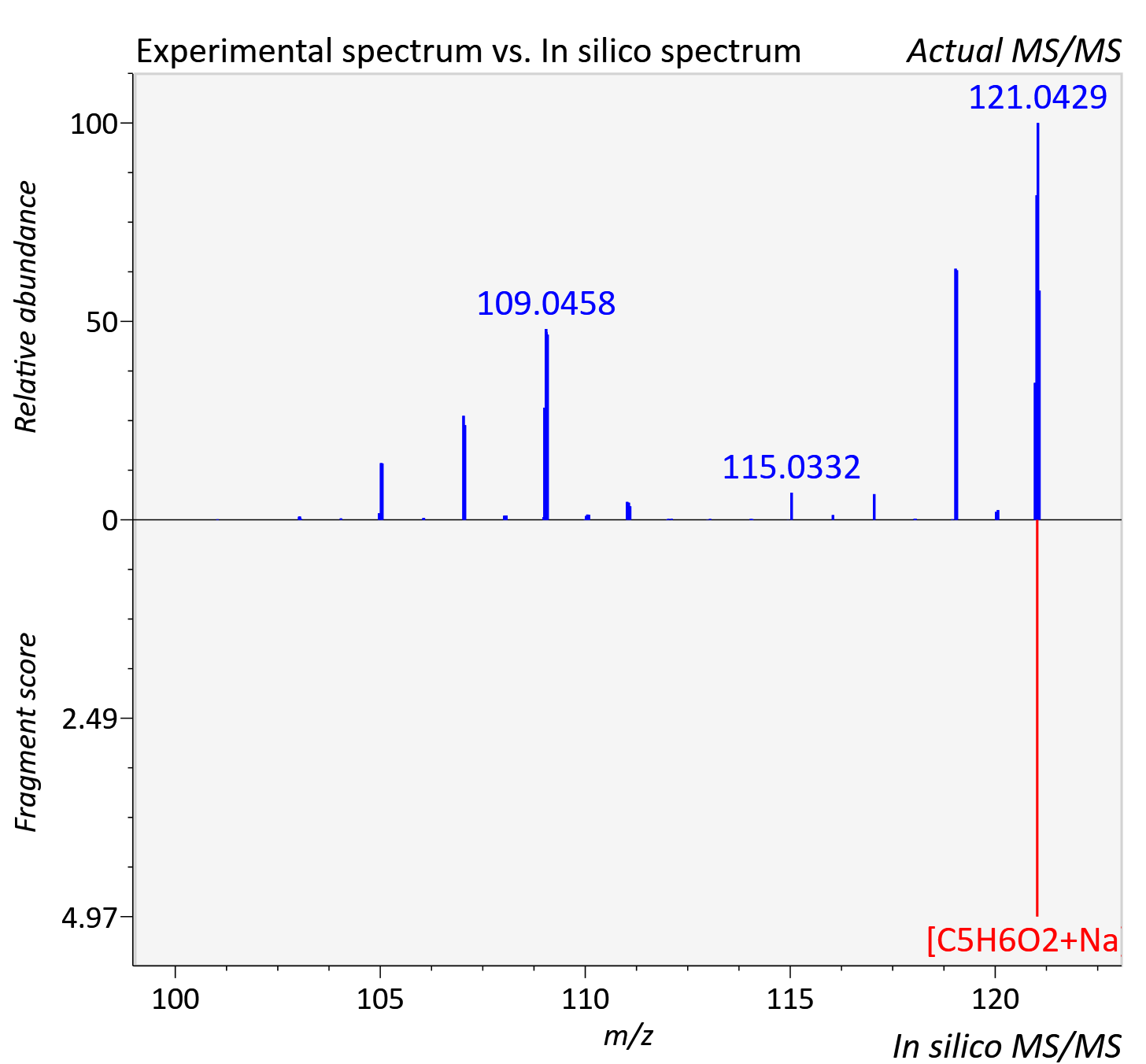
Experimental data
| Retention time | 17.92 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 121.026 | Theoretical mz | 121.026 |
| Error | 0.15 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.7999 |
Identifiers and class information
| Inchi key | XPFVYQJUAUNWIW-UHFFFAOYSA-N |
|---|---|
| Smiles | OCC=1OC=CC1 |
| Superclass | Organoheterocyclic compounds |
| Class | Heteroaromatic compounds |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T78198 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | P00491 | PNP |
| T78198 | DI0167 | Gout | [ICD-11: FA25] | P00491 | PNP |
| T78198 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P00491 | PNP |
| T78198 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P00491 | PNP |
| T78198 | DI0351 | Psoriasis | [ICD-11: EA90] | P00491 | PNP |
