Compound details
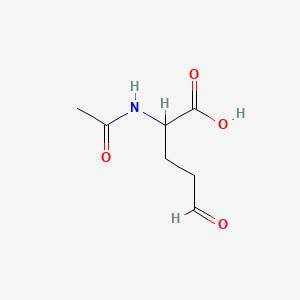
N-Acetyl-L-glutamate 5-semialdehyde
| Compound ID | CDAMM02904 |
|---|---|
| Common name | N-Acetyl-L-glutamate 5-semialdehyde | IUPAC name | 2-acetamido-5-oxopentanoic acid |
| Molecular formula | C7H11NO4 |
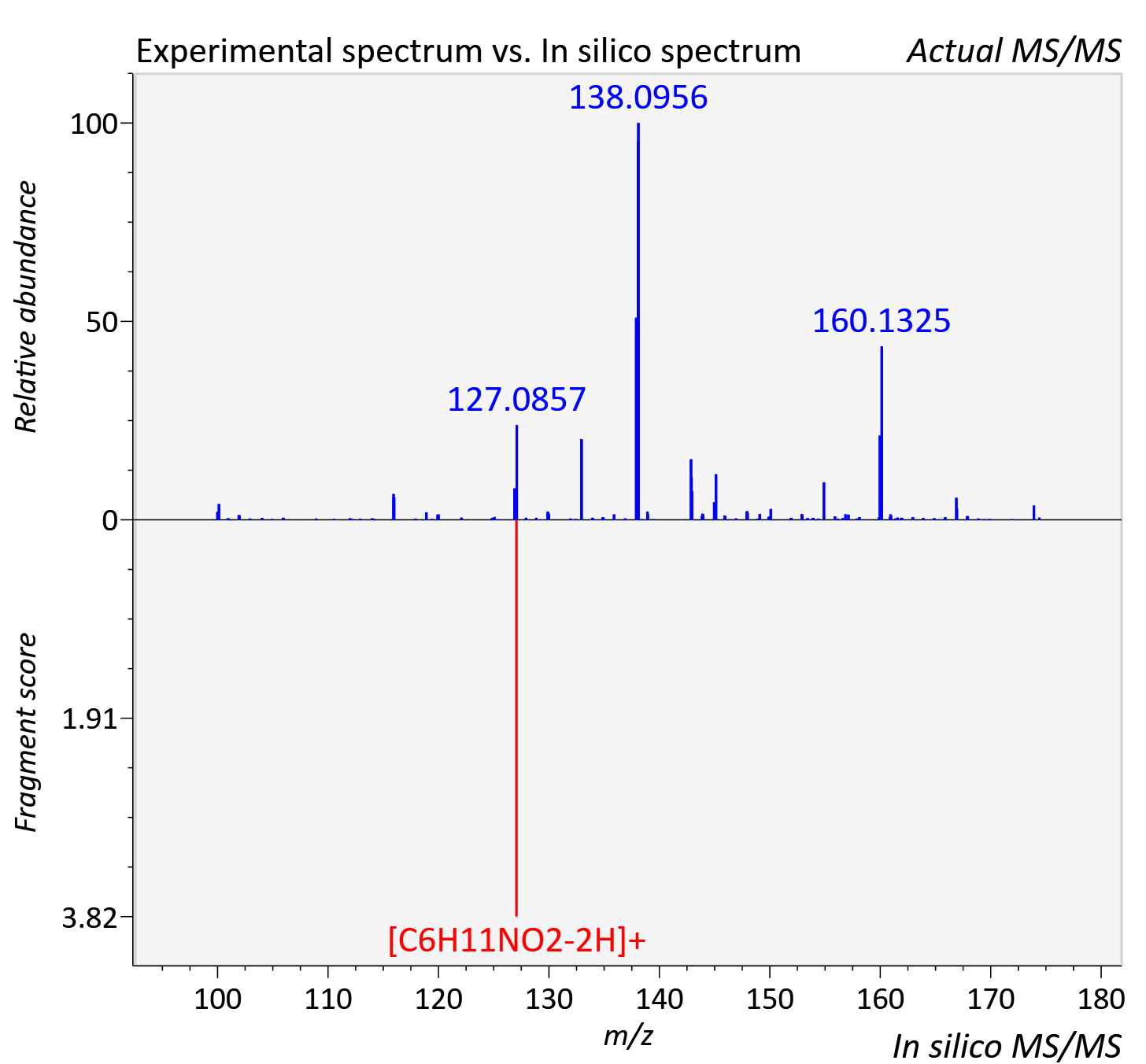
Experimental data
| Retention time | 0.51 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 174.074 | Theoretical mz | 174.076 |
| Error | 11.93 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.8546 |
Identifiers and class information
| Inchi key | BCPSFKBPHHBDAI-LURJTMIESA-N |
|---|---|
| Smiles | O=CCCC(NC(=O)C)C(=O)O |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P12821 | ACE | Angiotensin-converting enzyme | T98311 | SwissTargetPrediction |
| P00797 | REN | Renin | T61622 | SwissTargetPrediction |
| P09960 | LTA4H | Leukotriene A4 hydrolase | T03691 | SwissTargetPrediction |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SwissTargetPrediction |
| P08514 | ITGA2B | Integrin alpha-IIb/beta-3 | T50688 | SwissTargetPrediction |
| Q96RI1 | NR1H4 | Bile acid receptor FXR | T51426 | SwissTargetPrediction |
| P55210 | CASP7 | Caspase-7 | T72252 | SwissTargetPrediction |
| P08473 | MME | Neprilysin | T05409 | SwissTargetPrediction |
| P34913 | EPHX2 | Epoxide hydratase | T35734 | SwissTargetPrediction |
| P15121 | AKR1B1 | Aldose reductase | T26623 | SwissTargetPrediction |
| Q9H4A4 | RNPEP | Aminopeptidase B | T57818 | SwissTargetPrediction |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SwissTargetPrediction |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SwissTargetPrediction |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| O15229 | KMO | Kynurenine 3-monooxygenase | T50973 | SwissTargetPrediction |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SwissTargetPrediction |
| Q9Y5Y4 | PTGDR2 | G protein-coupled receptor 44 | T61722 | SwissTargetPrediction |
| P04439 | HLA-A | HLA class I histocompatibility antigen A-3 | T08985 | SwissTargetPrediction |
| Q9Y2K7 | KDM2A | Lysine-specific demethylase 2A | T71949 | SwissTargetPrediction |
| P42574 | CASP3 | Caspase-3 | T57943 | SwissTargetPrediction |
| Q6ZMT4 | KDM7A | Lysine-specific demethylase 7A/7B | T18784 | SwissTargetPrediction |
| P28074 | PSMB5 | Proteasome Macropain subunit MB1 | T49031 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SwissTargetPrediction and SEA |
| Q92820 | GGH | Gamma-glutamyl hydrolase | T71646 | SEA |
| Q9UPP1 | PHF8 | Histone lysine demethylase PHF8 | T00933 | SwissTargetPrediction |
| Q15393 | SF3B3 | Splicing factor 3B subunit 3 | T96723 | SEA |
| P09668 | CTSH | Cathepsin (H and K) | T40103 | SEA |
| O95551 | TDP2 | Tyrosyl-DNA phosphodiesterase 2 | T04696 | SwissTargetPrediction |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SEA |
| Q16348 | PEPT2 | Solute carrier family 15 member 2 | T52067 | SEA |
| Q9NXA8 | SIRT5 | NAD-dependent deacetylase sirtuin-5 | T91940 | SEA |
| Q9BX92 | CFB | Complement factor B | T66011 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T98311 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P12821 | ACE |
| T98311 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P12821 | ACE |
| T61622 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P00797 | REN |
| T03691 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P09960 | LTA4H |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T50688 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P08514 | ITGA2B |
| T50688 | DI0030 | Angina pectoris | [ICD-11: BA40] | P08514 | ITGA2B |
| T50688 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P08514 | ITGA2B |
| T51426 | DI0043 | Autoimmune liver disease | [ICD-11: DB96] | Q96RI1 | NR1H4 |
| T51426 | DI0077 | Cholelithiasis | [ICD-11: DC11] | Q96RI1 | NR1H4 |
| T51426 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | Q96RI1 | NR1H4 |
| T72252 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P55210 | CASP7 |
| T05409 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P08473 | MME |
| T35734 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P34913 | EPHX2 |
| T35734 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P34913 | EPHX2 |
| T26623 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P15121 | AKR1B1 |
| T26623 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P15121 | AKR1B1 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T97071 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | Q04609 | FOLH1 |
| T97071 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q04609 | FOLH1 |
| T61722 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | Q9Y5Y4 | PTGDR2 |
| T57943 | DI0060 | Brain cancer | [ICD-11: 2A00] | P42574 | CASP3 |
| T96723 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | Q15393 | SF3B3 |
| T96723 | DI0085 | Chronic myelomonocytic leukaemia | [ICD-11: 2A40] | Q15393 | SF3B3 |
| T96723 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | Q15393 | SF3B3 |
| T97071 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | Q04609 | FOLH1 |
| T97071 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q04609 | FOLH1 |
| T66011 | DI0170 | Haemolytic anemia | [ICD-11: 3A20-3A2Z] | Q9BX92 | CFB |
