Compound details
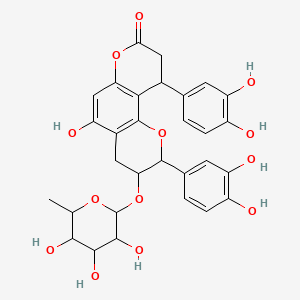
Glabraoside A
| Compound ID | CDAMM02874 |
|---|---|
| Common name | Glabraoside A | IUPAC name | 2,10-bis(3,4-dihydroxyphenyl)-5-hydroxy-3-(3,4,5-trihydroxy-6-methyloxan-2-yl)oxy-3,4,9,10-tetrahydro-2H-pyrano[2,3-f]chromen-8-one |
| Molecular formula | C30H30O13 |
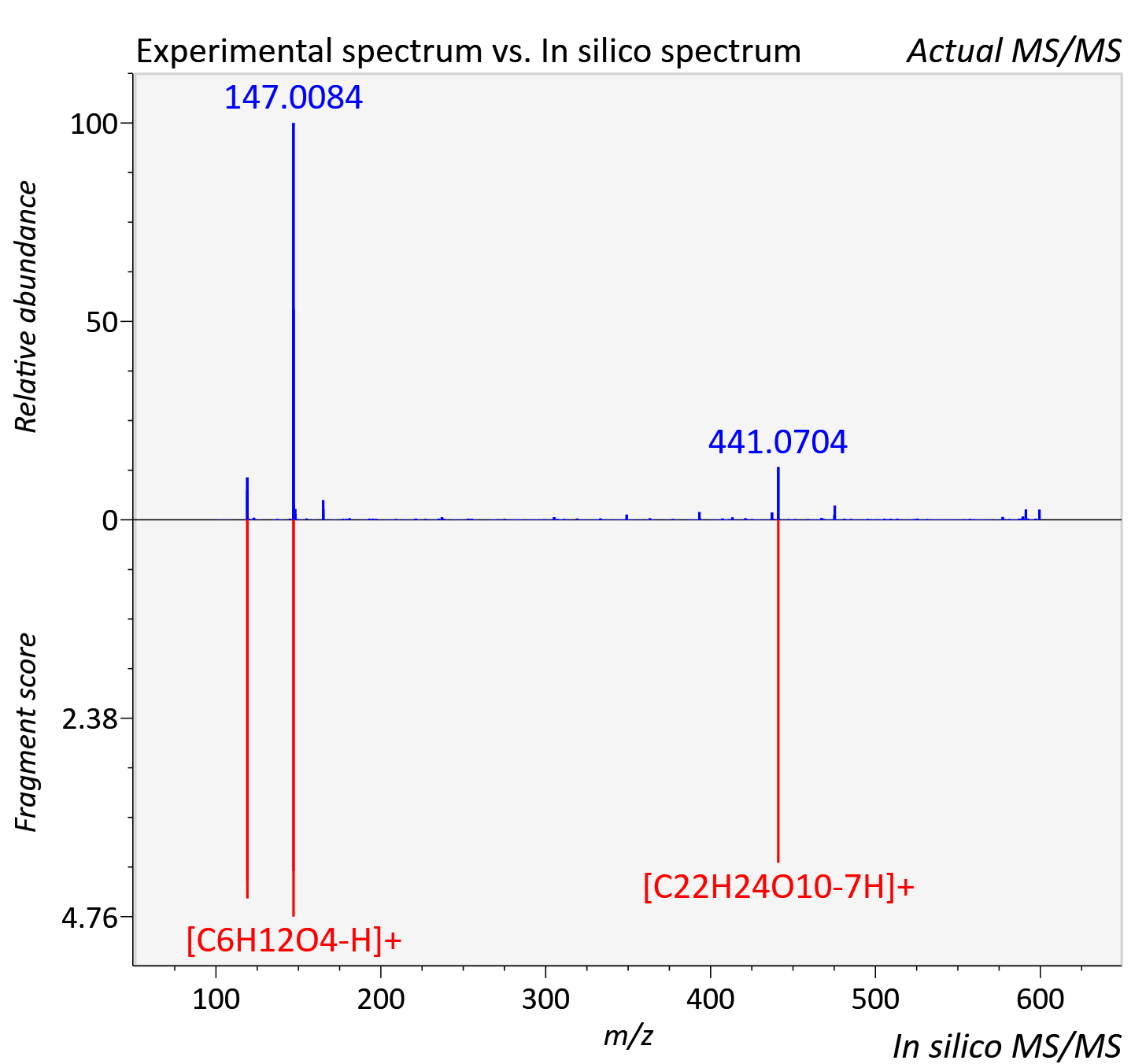
Experimental data
| Retention time | 15.44 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 599.168 | Theoretical mz | 599.176 |
| Error | 13.81 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.7004 |
Identifiers and class information
| Inchi key | OKKMHZTVGJIQAP-DWGIYUGRNA-N |
|---|---|
| Smiles | O=C1OC2=CC(O)=C3C(OC(C4=CC=C(O)C(O)=C4)C(OC5OC(C)C(O)C(O)C5O)C3)=C2C(C6=CC=C(O)C(O)=C6)C1 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q8IXJ6 | SIRT2 | NAD-dependent deacetylase sirtuin 2 | T83904 | SEA |
| P20151 | KLK2 | Kallikrein 2 | T01908 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P49763 | PGF | Placenta growth factor | T70792 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83904 | DI0365 | Retinopathy | [ICD-11: 9B71] | Q8IXJ6 | SIRT2 |
| T01908 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | P20151 | KLK2 |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T70792 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P49763 | PGF |
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
