Compound details
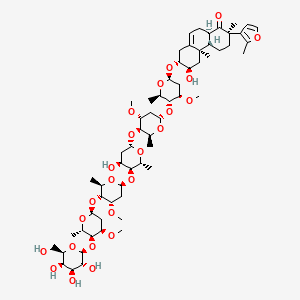
Sublanceoside L6
| Compound ID | CDAMM02870 |
|---|---|
| Common name | Sublanceoside L6 | IUPAC name | (2R,4aS,4bR,6R,7R,10aR)-6-hydroxy-7-[(2S,4S,5R,6R)-5-[(2S,4R,5R,6S)-5-[(2S,4S,5S,6R)-4-hydroxy-5-[(2S,4S,5R,6R)-4-methoxy-5-[(2S,4R,5S,6S)-4-methoxy-6-methyl-5-[(2S,3R,4S,5R,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxyoxan-2-yl]oxy-6-methyloxan-2-yl]oxy-6-methyloxan-2-yl]oxy-4-methoxy-6-methyloxan-2-yl]oxy-4-methoxy-6-methyloxan-2-yl]oxy-2,4b-dimethyl-2-(2-methylfuran-3-yl)-4,4a,5,6,7,8,10,10a-octahydro-3H-phenanthren-1-one |
| Molecular formula | C61H96O24 |
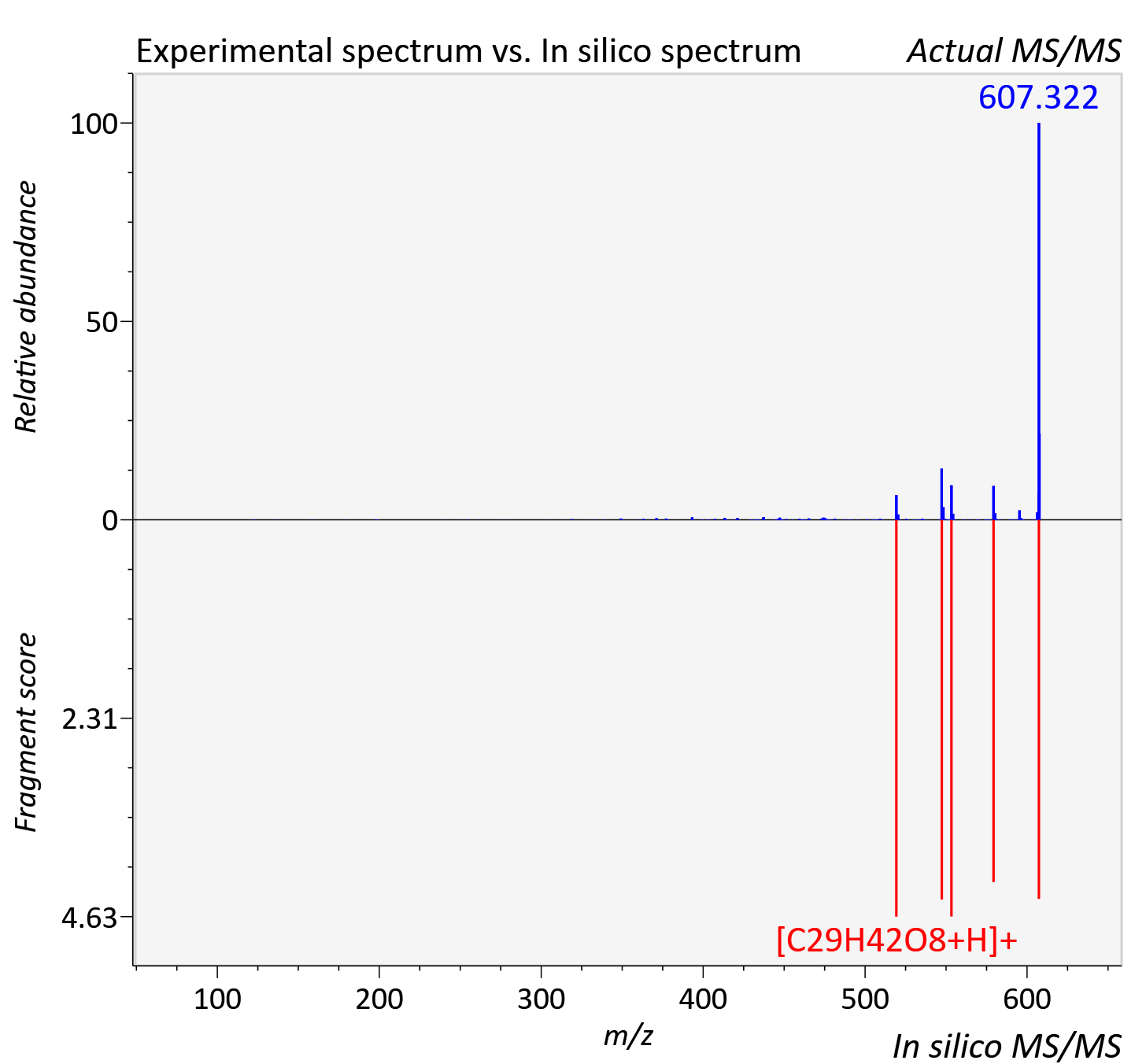
Experimental data
| Retention time | 11.78 |
|---|---|
| Adduct | [M+2H]2+ |
| Actual mz | 607.321 | Theoretical mz | 607.322 |
| Error | 2.76 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0034 |
Identifiers and class information
| Inchi key | RLGGUDAQMBDXLW-AEBBZHNZNA-N |
|---|---|
| Smiles | O=C1C2CC=C3CC(OC4OC(C)C(OC5OC(C)C(OC6OC(C)C(OC7OC(C)C(OC8OC(C)C(OC9OC(CO)C(O)C(O)C9O)C(OC)C8)C(OC)C7)C(O)C6)C(OC)C5)C(OC)C4)C(O)CC3(C)C2CCC1(C=%10C=COC%10C)C |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T86918 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P04746 | AMY2A |
| T86918 | DI0328 | Pancreatic malfunction | [ICD-11: DC30-DC3Z] | P04746 | AMY2A |
