Compound details
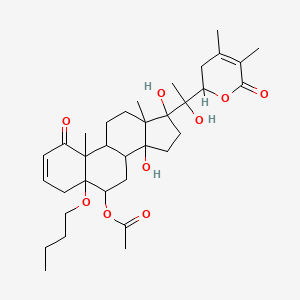
Physapruin B
| Compound ID | CDAMM02852 |
|---|---|
| Common name | Physapruin B | IUPAC name | [5-butoxy-17-[1-(4,5-dimethyl-6-oxo-2,3-dihydropyran-2-yl)-1-hydroxyethyl]-14,17-dihydroxy-10,13-dimethyl-1-oxo-6,7,8,9,11,12,15,16-octahydro-4H-cyclopenta[a]phenanthren-6-yl] acetate |
| Molecular formula | C34H50O9 |
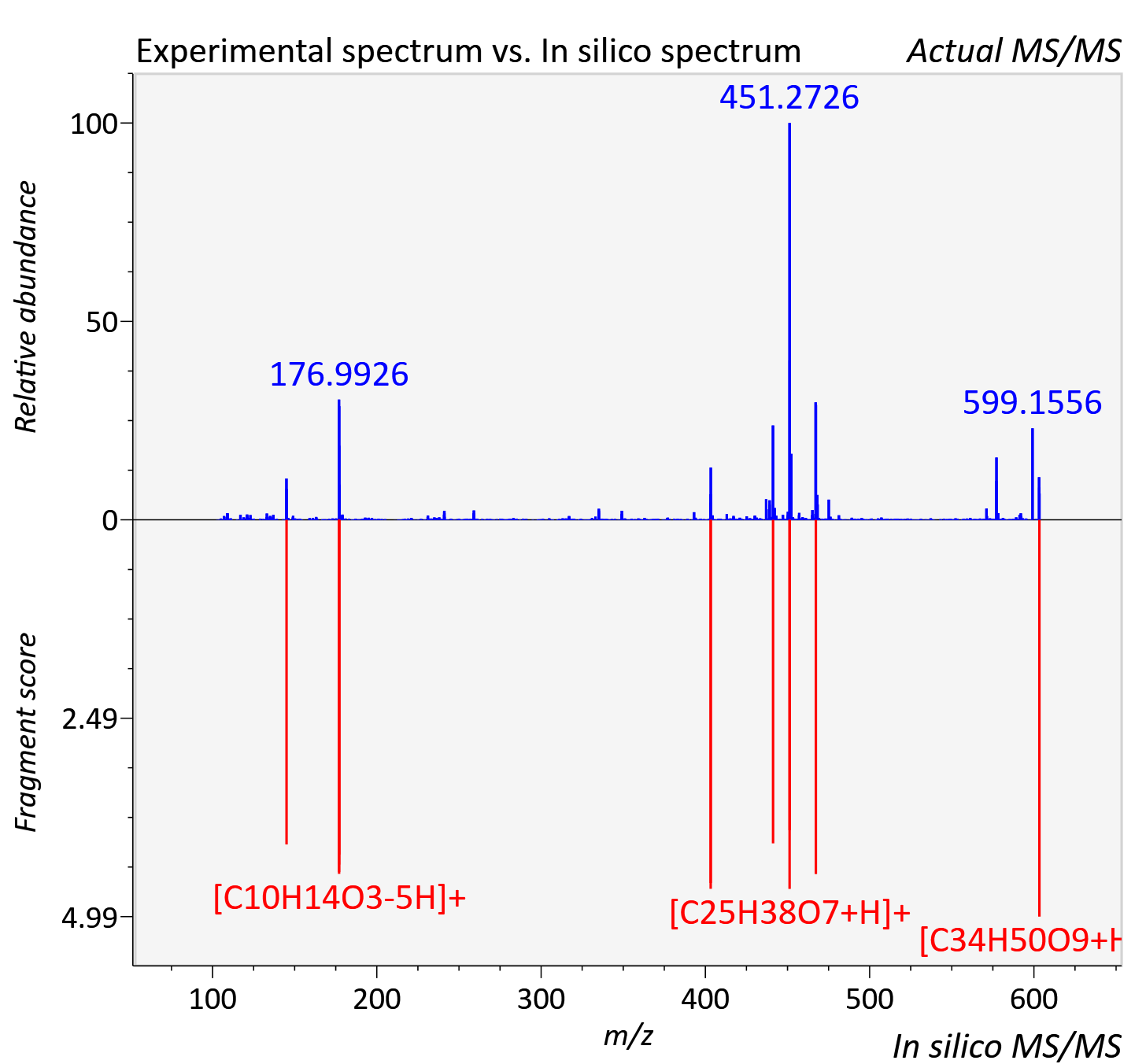
Experimental data
| Retention time | 15.56 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 603.355 | Theoretical mz | 603.352 |
| Error | 5.01 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.5429 |
Identifiers and class information
| Inchi key | BUTLOLBCWDNVGA-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC(CC(=C1C)C)C(O)(C)C2(O)CCC3(O)C4CC(OC(=O)C)C5(OCCCC)CC=CC(=O)C5(C)C4CCC32C |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SwissTargetPrediction |
| Q08499 | PDE4D | Phosphodiesterase 4D | T02001 | SwissTargetPrediction |
| Q02156 | PRKCE | Protein kinase C epsilon | T00895 | SwissTargetPrediction |
| Q05655 | PRKCD | Protein kinase C delta | T44861 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T02001 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | Q08499 | PDE4D |
| T00895 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | Q02156 | PRKCE |
| T00895 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q02156 | PRKCE |
| T44861 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | Q05655 | PRKCD |
