Compound details
Grewinol
| Compound ID | CDAMM02834 |
|---|---|
| Common name | Grewinol | IUPAC name | 22-hydroxytetratriacontan-13-one |
| Molecular formula | C34H68O2 |
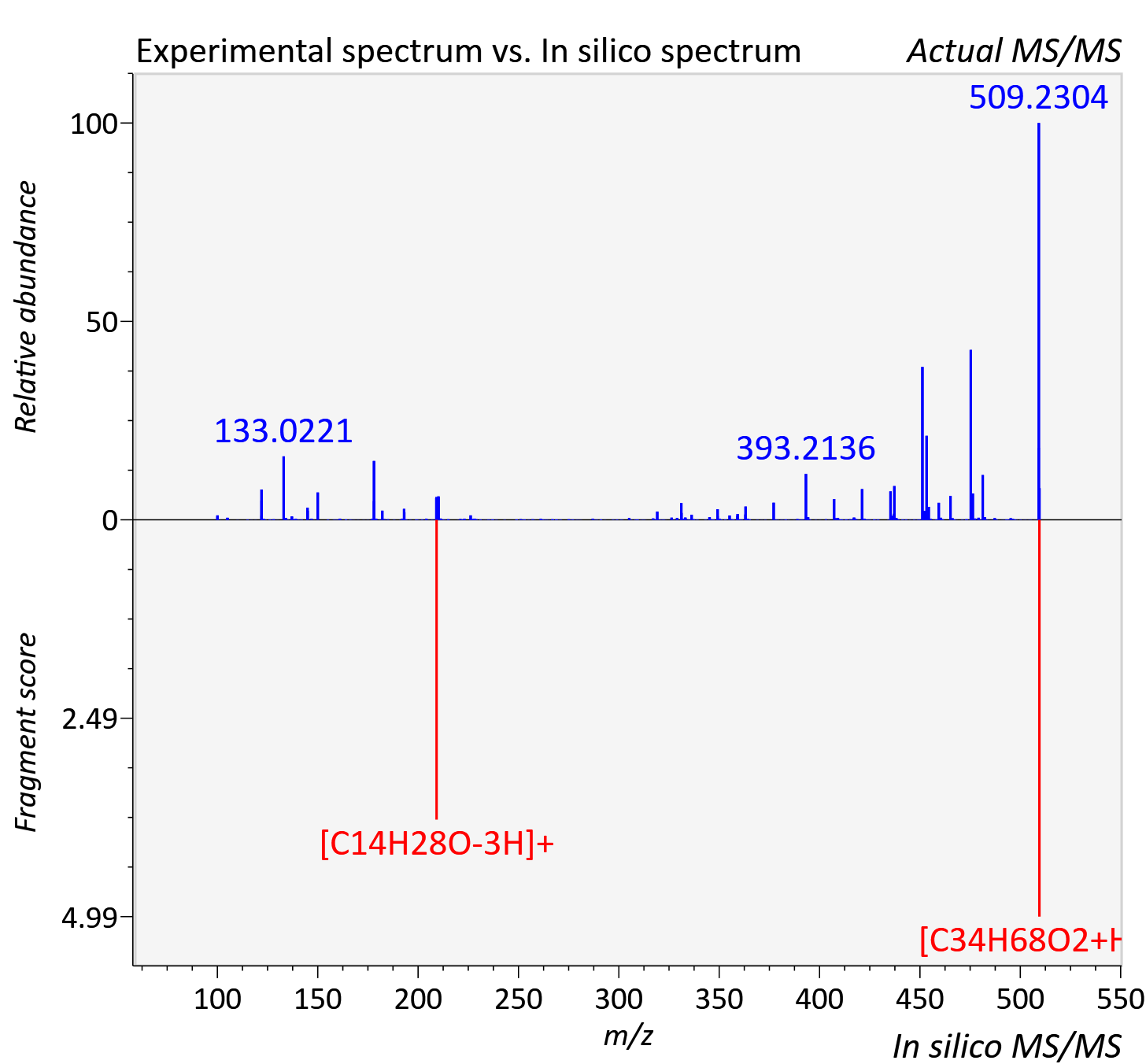
Experimental data
| Retention time | 3.91 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 509.522 | Theoretical mz | 509.529 |
| Error | 13.3 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.0819 |
Identifiers and class information
| Inchi key | LJBLOIRMOMZGOI-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(CCCCCCCCCCCC)CCCCCCCCC(O)CCCCCCCCCCCC |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SEA |
| P04054 | PLA2G1B | Phospholipase A2 group 1B | T31479 | SEA |
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SEA |
| P34913 | EPHX2 | Epoxide hydratase | T35734 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| Q07075 | ENPEP | Aminopeptidase A | T31956 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| P08254 | MMP3 | Matrix metalloproteinase 3 | T86702 | SEA |
| P43116 | PTGER2 | Prostanoid EP2 receptor | T38529 | SEA |
| P04035 | HMGCR | HMG-CoA reductase | T53585 | SEA |
| Q15722 | LTB4R | Leukotriene B4 receptor 1 | T59626 | SEA |
| P08263 | GSTA1 | Glutathione S-transferase A1 | T17221 | SEA |
| P35408 | PTGER4 | Prostanoid EP4 receptor | T18876 | SEA |
| P0DJD9 | PGA5 | Pepsin A | T17863 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q9Y4D2 | DAGLA | Sn1-specific diacylglycerol lipase alpha | T03150 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P40189 | IL6ST | Interleukin-6 receptor subunit beta | T93440 | SEA |
| Q13822 | ENPP2 | Autotaxin | T63512 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9H1C0 | LPAR5 | Lysophosphatidic acid receptor 5 | T18616 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| O95977 | S1PR4 | Sphingosine 1-phosphate receptor Edg-6 | T17523 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| Q9BXC1 | GPR174 | Probable G-protein coupled receptor 174 | T87661 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P40394 | ADH7 | Alcohol dehydrogenase class IV | T09770 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T31479 | DI0230 | Leishmaniasis | [ICD-11: 1F54] | P04054 | PLA2G1B |
| T31479 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P04054 | PLA2G1B |
| T31479 | DI0367 | Rosacea | [ICD-11: ED90] | P04054 | PLA2G1B |
| T31479 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P04054 | PLA2G1B |
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T35734 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P34913 | EPHX2 |
| T35734 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P34913 | EPHX2 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T31956 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q07075 | ENPEP |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T86702 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P08254 | MMP3 |
| T86702 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P08254 | MMP3 |
| T86702 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | P08254 | MMP3 |
| T86702 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P08254 | MMP3 |
| T86702 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P08254 | MMP3 |
| T86702 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P08254 | MMP3 |
| T86702 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P08254 | MMP3 |
| T38529 | DI0003 | Abortion | [ICD-11: JA00] | P43116 | PTGER2 |
| T38529 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43116 | PTGER2 |
| T38529 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P43116 | PTGER2 |
| T38529 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43116 | PTGER2 |
| T38529 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43116 | PTGER2 |
| T38529 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P43116 | PTGER2 |
| T38529 | DI0413 | Transplant rejection | [ICD-11: NE84] | P43116 | PTGER2 |
| T53585 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P04035 | HMGCR |
| T53585 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P04035 | HMGCR |
| T53585 | DI0128 | Dyslipidemia | [ICD-11: 5C80-5C81] | P04035 | HMGCR |
| T53585 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P04035 | HMGCR |
| T53585 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P04035 | HMGCR |
| T53585 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P04035 | HMGCR |
| T53585 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P04035 | HMGCR |
| T59626 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | Q15722 | LTB4R |
| T59626 | DI0355 | Pulmonary disease | [ICD-11: 1B10-1F85] | Q15722 | LTB4R |
| T59626 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | Q15722 | LTB4R |
| T18876 | DI0037 | Asthma | [ICD-11: CA23] | P35408 | PTGER4 |
| T18876 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P35408 | PTGER4 |
| T18876 | DI0264 | Migraine | [ICD-11: 8A80] | P35408 | PTGER4 |
| T18876 | DI0303 | Non-small-cell lung cancer | [ICD-11: 2C25] | P35408 | PTGER4 |
| T18876 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35408 | PTGER4 |
| T18876 | DI0360 | Rectum cancer | [ICD-11: 2B92] | P35408 | PTGER4 |
| T18876 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35408 | PTGER4 |
| T18876 | DI0414 | Transplanted organ/tissue | [ICD-11: QB63] | P35408 | PTGER4 |
| T93440 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P40189 | IL6ST |
| T63512 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q13822 | ENPP2 |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
