Compound details
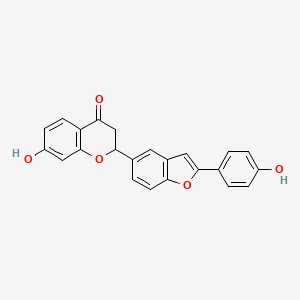
Lophirone I
| Compound ID | CDAMM02764 |
|---|---|
| Common name | Lophirone I | IUPAC name | 7-hydroxy-2-[2-(4-hydroxyphenyl)-1-benzofuran-5-yl]-2,3-dihydrochromen-4-one |
| Molecular formula | C23H16O5 |
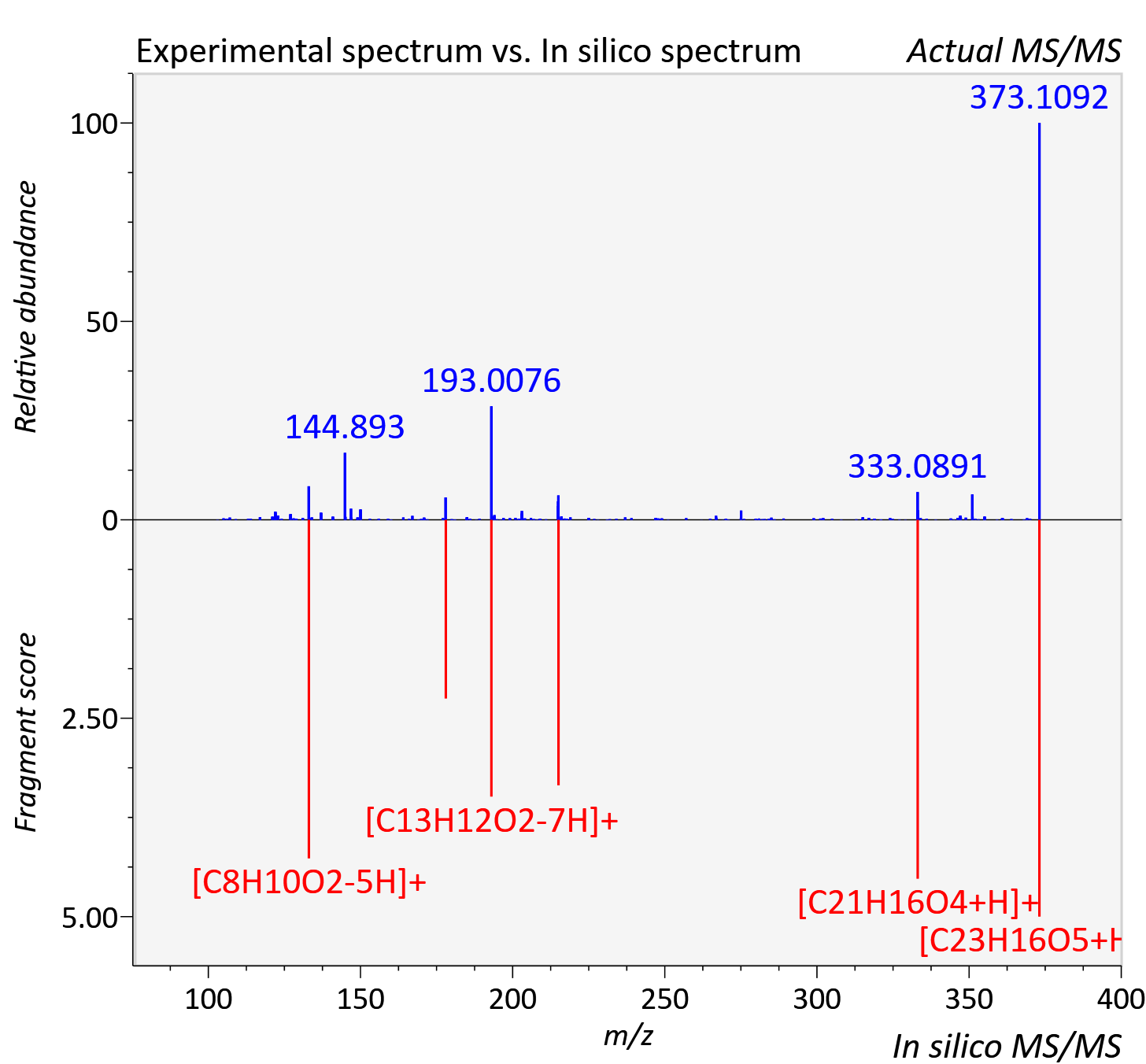
Experimental data
| Retention time | 1.76 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 373.107 | Theoretical mz | 373.107 |
| Error | 0.74 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0372 |
Identifiers and class information
| Inchi key | FKAOXSDPCYTXNP-ANBDAQEENA-N |
|---|---|
| Smiles | O=C1C2=CC=C(O)C=C2OC(C3=CC=C4OC(=CC4=C3)C=5C=CC(O)=CC5)C1 |
| Superclass | Phenylpropanoids and polyketides |
| Class | 2-arylbenzofuran flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P05067 | APP | Beta amyloid A4 protein | T87024 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| P11086 | PNMT | Phenylethanolamine N-methyltransferase | T56496 | SwissTargetPrediction |
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SEA |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SEA |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SEA |
| P14061 | HSD17B1 | Estradiol 17-beta-dehydrogenase 1 | T44011 | SwissTargetPrediction |
| P56817 | BACE1 | Beta-secretase 1 | T79031 | SwissTargetPrediction |
| O14965 | AURKA | Serine/threonine-protein kinase Aurora-A | T87675 | SwissTargetPrediction |
| P20151 | KLK2 | Kallikrein 2 | T01908 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P49763 | PGF | Placenta growth factor | T70792 | SEA |
| P47989 | XDH | Xanthine dehydrogenase | T40954 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SwissTargetPrediction and SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SwissTargetPrediction and SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| P27169 | PON1 | Serum paraoxonase/arylesterase 1 | T34562 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
| Q96HK3 | CALM | Calmodulin | T39610 | SEA |
| Q9UK17 | KCND3 | Voltage-gated potassium channel Kv4.3 | T74500 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T87024 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P05067 | APP |
| T87024 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P05067 | APP |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T79031 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P56817 | BACE1 |
| T87675 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | O14965 | AURKA |
| T87675 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | O14965 | AURKA |
| T01908 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | P20151 | KLK2 |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T70792 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P49763 | PGF |
| T40954 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | P47989 | XDH |
| T40954 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P47989 | XDH |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
| T39610 | DI0068 | Cardiac arrhythmia | [ICD-11: BC9Z] | Q96HK3 | CALM |
| T39610 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q96HK3 | CALM |
| T39610 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q96HK3 | CALM |
| T74500 | DI0004 | Acidosis | [ICD-11: 5C73] | Q9UK17 | KCND3 |
