Compound details
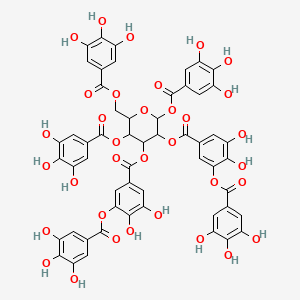
2,3-O-digalloyl-1,4,6-tri-O-beta-D-galloylglucose
| Compound ID | CDAMM02758 |
|---|---|
| Common name | 2,3-O-digalloyl-1,4,6-tri-O-beta-D-galloylglucose | IUPAC name | [4,5-bis[[3,4-dihydroxy-5-(3,4,5-trihydroxybenzoyl)oxybenzoyl]oxy]-3,6-bis[(3,4,5-trihydroxybenzoyl)oxy]oxan-2-yl]methyl 3,4,5-trihydroxybenzoate |
| Molecular formula | C55H40O34 |
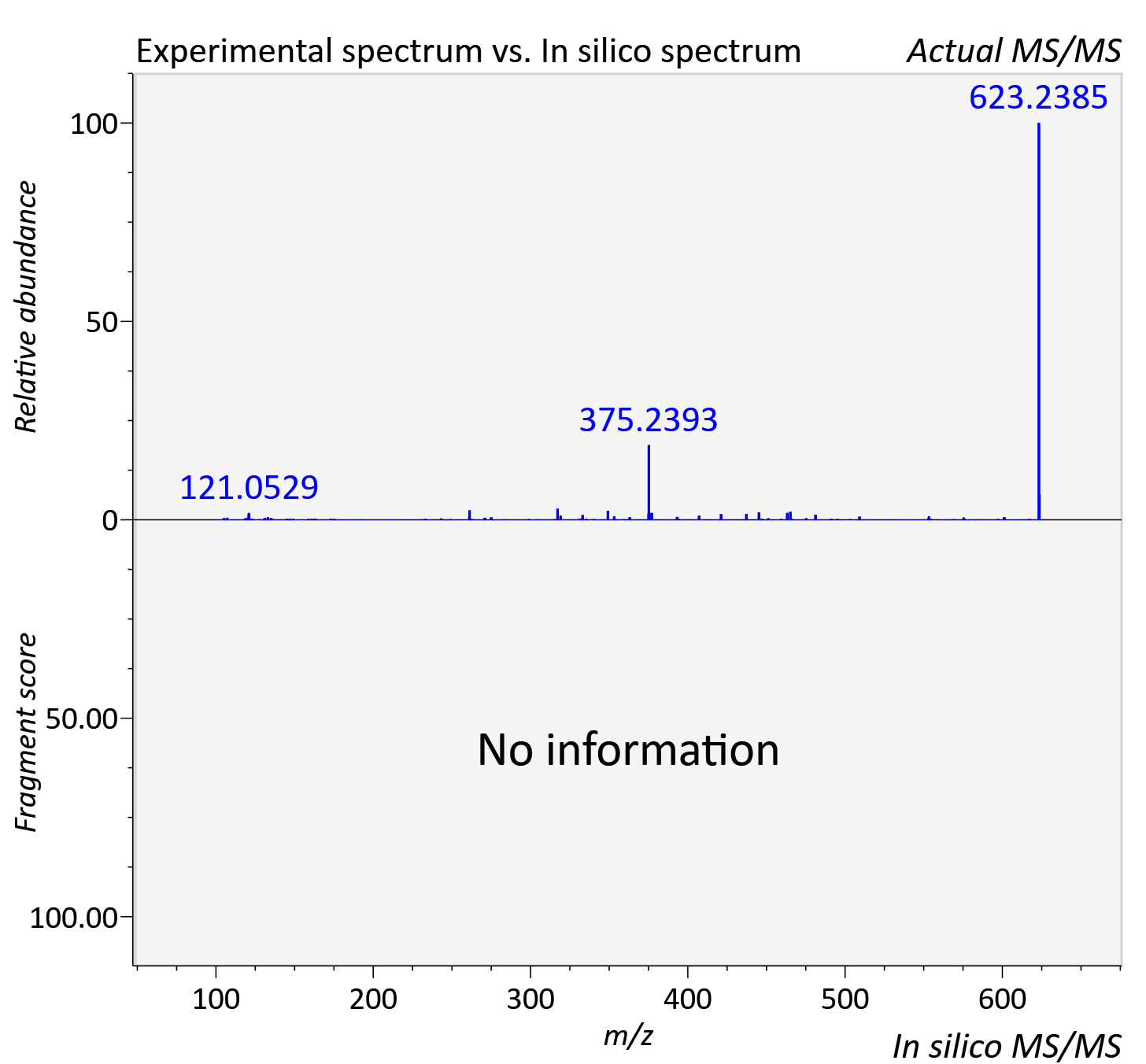
Experimental data
| Retention time | 9.62 |
|---|---|
| Adduct | [M+2H]2+ |
| Actual mz | 623.08 | Theoretical mz | 623.077 |
| Error | 3.57 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.3106 |
Identifiers and class information
| Inchi key | CWFXJAOBRIWZTE-YIVDHKKMSA-N |
|---|---|
| Smiles | O=C(OC1=CC(=CC(O)=C1O)C(=O)OC2C(OC(=O)C3=CC(O)=C(O)C(O)=C3)OC(COC(=O)C4=CC(O)=C(O)C(O)=C4)C(OC(=O)C5=CC(O)=C(O)C(O)=C5)C2OC(=O)C6=CC(O)=C(O)C(OC(=O)C7=CC(O)=C(O)C(O)=C7)=C6)C8=CC(O)=C(O)C(O)=C8 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Tannins |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SEA |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SEA |
| P00915 | CA1 | Carbonic anhydrase I | T13201 | SEA |
| P00918 | CA2 | Carbonic anhydrase II | T20401 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| P05121 | SERPINE1 | Plasminogen activator inhibitor-1 | T15556 | SEA |
| P14151 | SELL | Leukocyte adhesion molecule-1 | T60526 | SEA |
| Q9NUW8 | TDP1 | Tyrosyl-DNA phosphodiesterase 1 | T33492 | SEA |
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T13201 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00915 | CA1 |
| T13201 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00915 | CA1 |
| T20401 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P00918 | CA2 |
| T20401 | DI0137 | Essential hypertension | [ICD-11: BA00] | P00918 | CA2 |
| T20401 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00918 | CA2 |
| T20401 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P00918 | CA2 |
| T20401 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P00918 | CA2 |
| T20401 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00918 | CA2 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T15556 | DI0037 | Asthma | [ICD-11: CA23] | P05121 | SERPINE1 |
| T15556 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | P05121 | SERPINE1 |
| T60526 | DI0037 | Asthma | [ICD-11: CA23] | P14151 | SELL |
| T60526 | DI0088 | Circulatory system disease | [ICD-11: BE2Z] | P14151 | SELL |
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
