Compound details
Longipedlactone J
| Compound ID | CDAMM02756 |
|---|---|
| Common name | Longipedlactone J | IUPAC name | [1-hydroxy-8,8,13-trimethyl-17-methylidene-16-[1-(5-methyl-6-oxo-2,3-dihydropyran-2-yl)ethyl]-6-oxo-7-oxatetracyclo[10.7.0.03,9.013,18]nonadeca-2,4,15-trien-10-yl] acetate |
| Molecular formula | C32H40O7 |
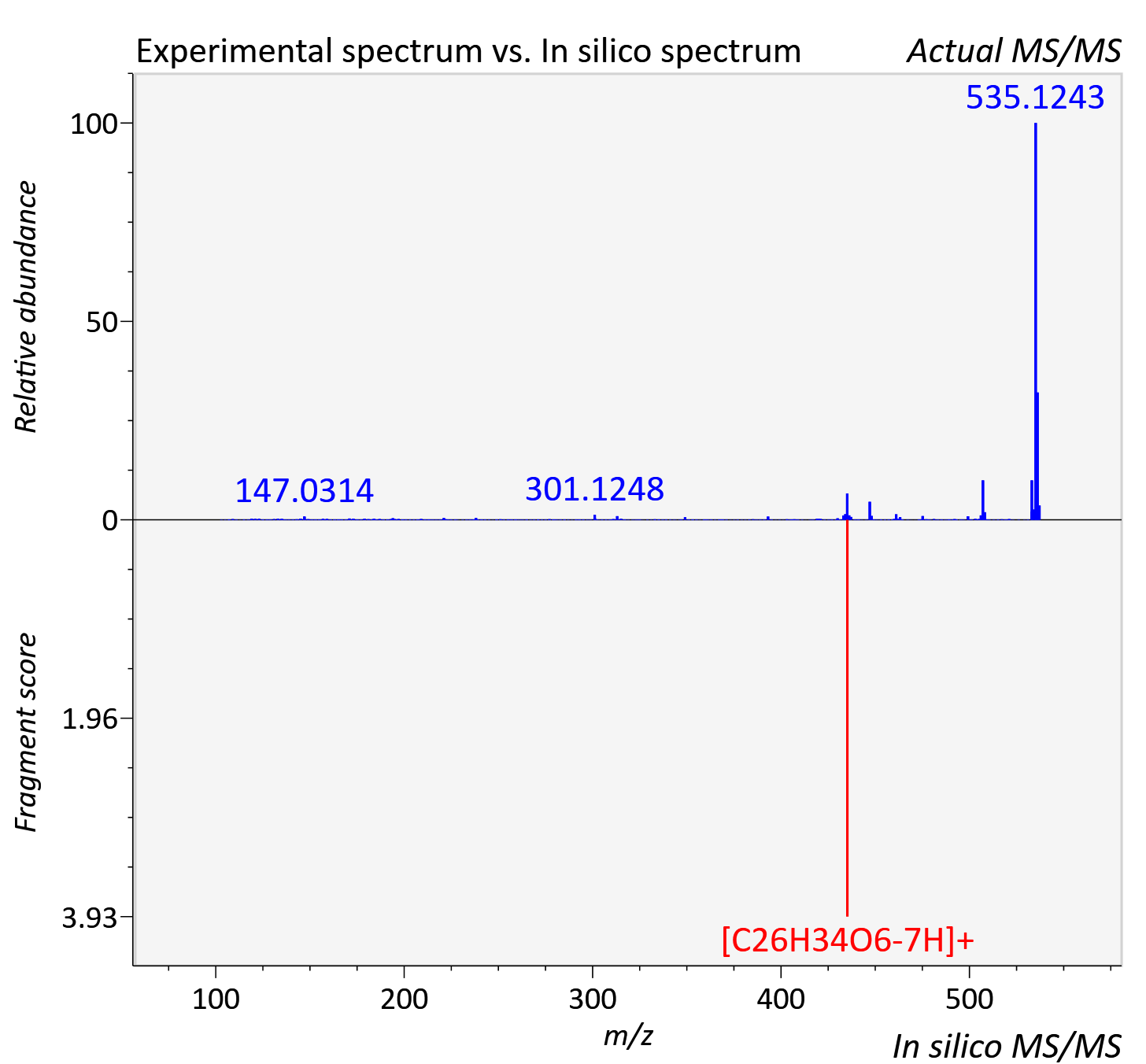
Experimental data
| Retention time | 13.56 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 537.29 | Theoretical mz | 537.284 |
| Error | 9.91 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.5033 |
Identifiers and class information
| Inchi key | VPYOIPWGVNGDBV-UKTXQOCXNA-N |
|---|---|
| Smiles | O=C1OC(C)(C)C2C(C=C1)=CC3(O)CC4C(=C)C(=CCC4(C)C3CC2OC(=O)C)C(C)C5OC(=O)C(=CC5)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|
