Compound details
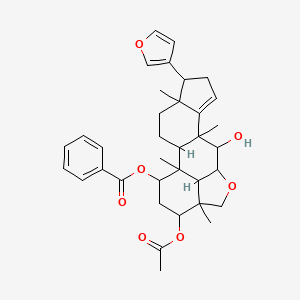
Meliavolkinin
| Compound ID | CDAMM02730 |
|---|---|
| Common name | Meliavolkinin | IUPAC name | [16-acetyloxy-6-(furan-3-yl)-11-hydroxy-1,5,10,15-tetramethyl-13-oxapentacyclo[10.6.1.02,10.05,9.015,19]nonadec-8-en-18-yl] benzoate |
| Molecular formula | C35H42O7 |
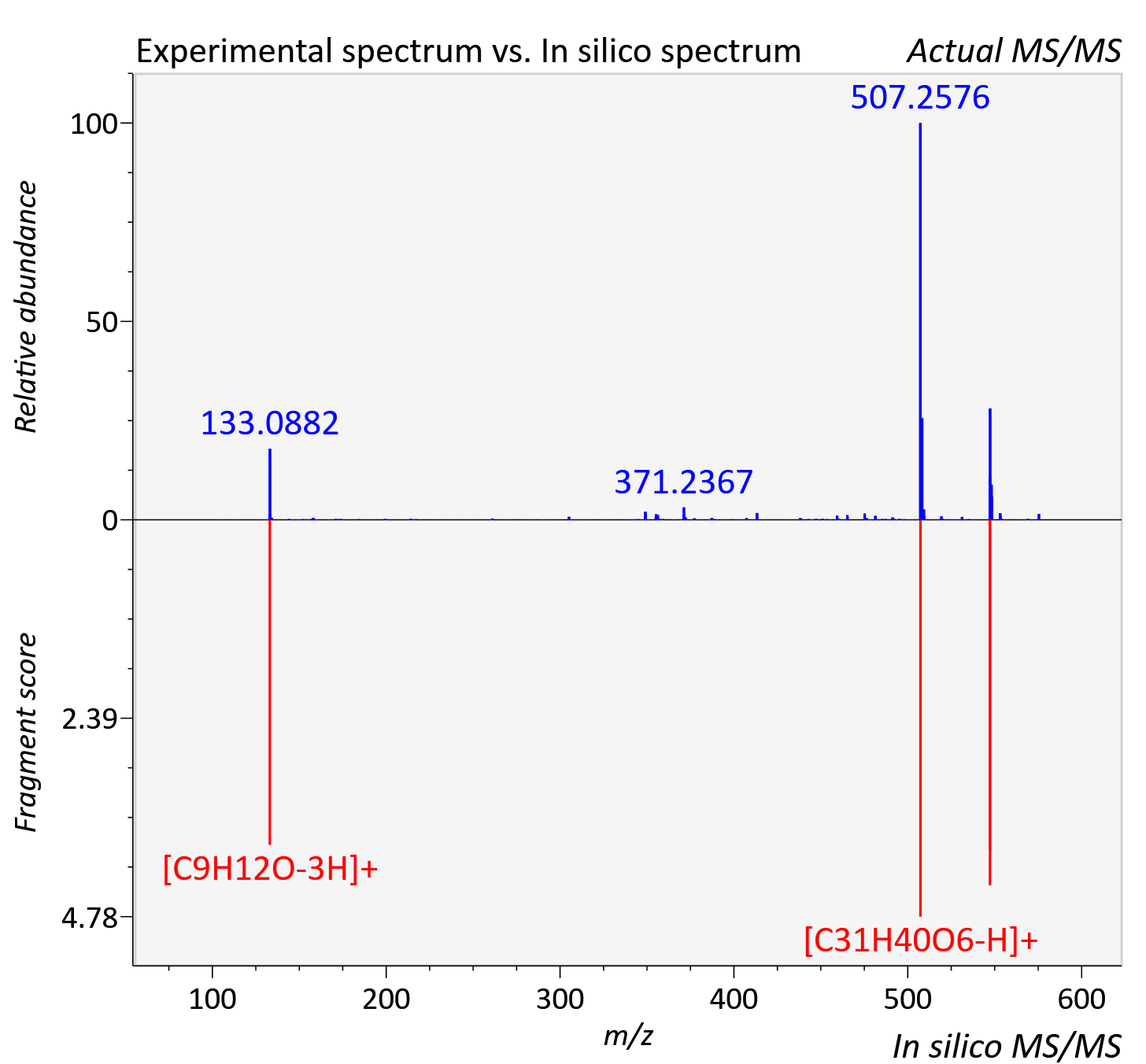
Experimental data
| Retention time | 11.99 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 575.308 | Theoretical mz | 575.3 |
| Error | 14.29 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.9714 |
Identifiers and class information
| Inchi key | OVRHPVPNJONXSX-JNMWFZIXNA-N |
|---|---|
| Smiles | O=C(OC1CC(OC(=O)C)C2(C)COC3C(O)C4(C5=CCC(C6=COC=C6)C5(C)CCC4C1(C)C32)C)C=7C=CC=CC7 |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P41145 | OPRK1 | Kappa Opioid receptor | T60693 | SEA |
| P07900 | HSP90AA1 | Heat shock protein HSP 90-alpha | T18477 | SwissTargetPrediction |
| O75908 | SOAT2 | Acyl coenzyme A:cholesterol acyltransferase 2 | T14463 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T60693 | DI0304 | Non-specific cutaneous vascular symptom | [ICD-11: ME64] | P41145 | OPRK1 |
| T60693 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P41145 | OPRK1 |
| T60693 | DI0349 | Pruritus | [ICD-11: EC90] | P41145 | OPRK1 |
| T18477 | DI0015 | Acute upper respiratory infection | [ICD-11: CA07] | P07900 | HSP90AA1 |
| T18477 | DI0037 | Asthma | [ICD-11: CA23] | P07900 | HSP90AA1 |
| T14463 | DI0017 | Adrenal cancer | [ICD-11: 2D11] | O75908 | SOAT2 |
