Compound details
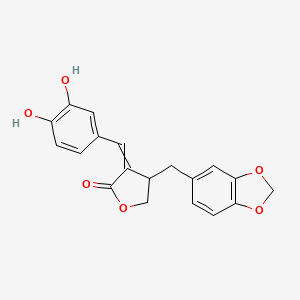
Piperphilippinin IV
| Compound ID | CDAMM02726 |
|---|---|
| Common name | Piperphilippinin IV | IUPAC name | 4-(1,3-benzodioxol-5-ylmethyl)-3-[(3,4-dihydroxyphenyl)methylidene]oxolan-2-one |
| Molecular formula | C19H16O6 |
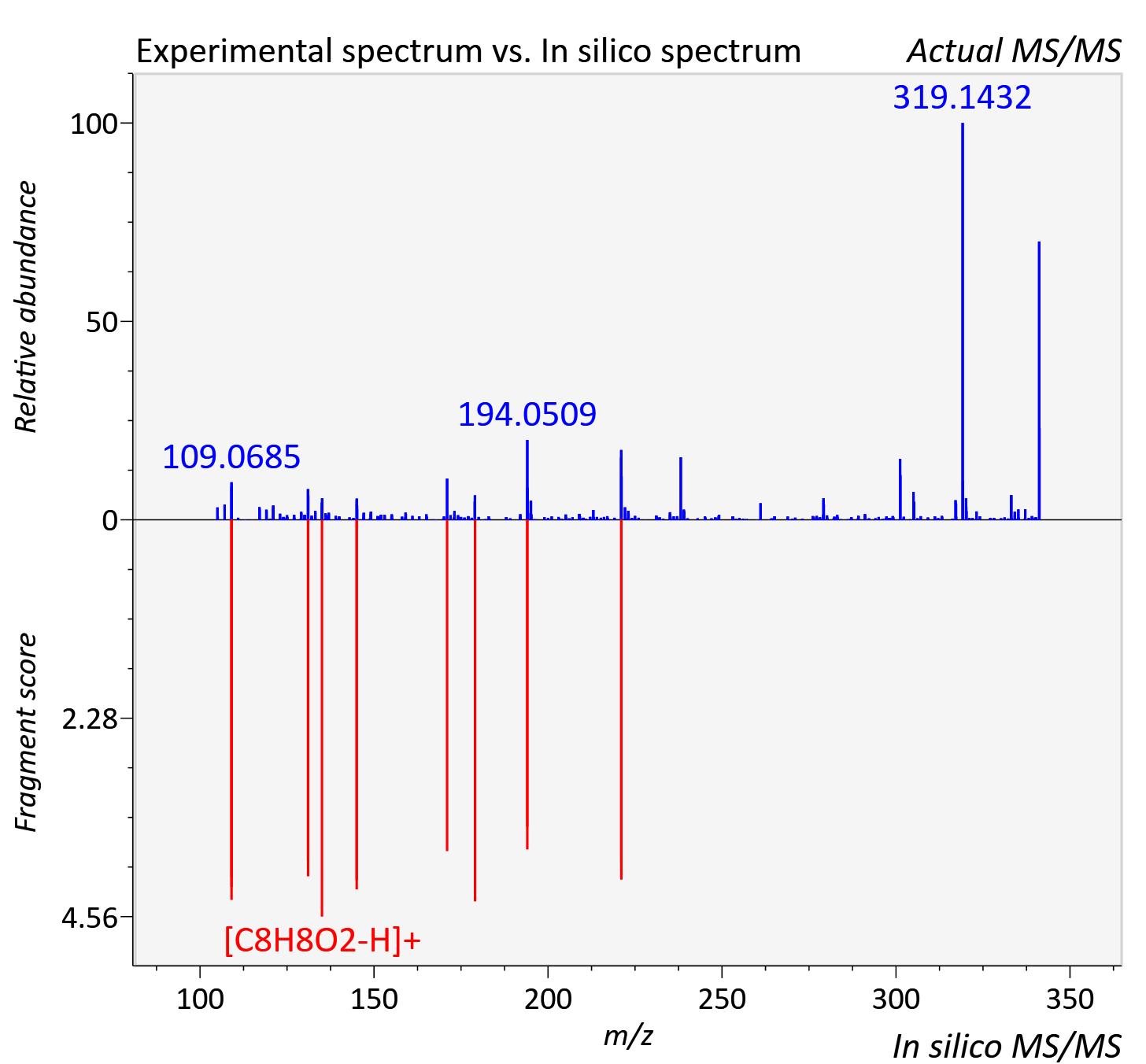
Experimental data
| Retention time | 10.15 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 341.105 | Theoretical mz | 341.102 |
| Error | 6.81 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.929 |
Identifiers and class information
| Inchi key | GWOSVBIYCRFNFK-USSUKOEXNA-N |
|---|---|
| Smiles | O=C1OCC(C1=CC2=CC=C(O)C(O)=C2)CC3=CC=C4OCOC4=C3 |
| Superclass | Lignans, neolignans and related compounds |
| Class | Furanoid lignans |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| Q15078 | CDK5R1 | Cyclin-dependent kinase 5/CDK5 activator 1 | T09513 | SEA |
| Q00535 | CDK5 | Cyclin-dependent kinase 5/CDK5 activator 1 | T20973 | SEA |
| P17936 | IGFBP3 | Insulin-like growth factor binding protein 3 | T33455 | SEA |
| P01130 | LDLR | LDL receptor | T94692 | SEA |
| Q92630 | DYRK2 | Dual-specificity tyrosine-phosphorylation regulated kinase 2 | T39486 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T20973 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | Q00535 | CDK5 |
| T33455 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P17936 | IGFBP3 |
| T94692 | DI0238 | Lung cancer | [ICD-11: 2C25] | P01130 | LDLR |
