Compound details
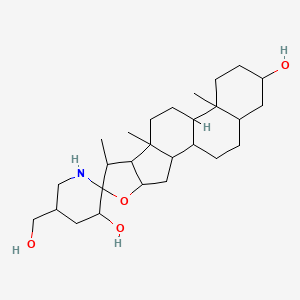
Isoesculeogenin A
| Compound ID | CDAMM02702 |
|---|---|
| Common name | Isoesculeogenin A | IUPAC name | 5\'-(hydroxymethyl)-7,9,13-trimethylspiro[5-oxapentacyclo[10.8.0.02,9.04,8.013,18]icosane-6,2\'-piperidine]-3\',16-diol |
| Molecular formula | C27H45NO4 |
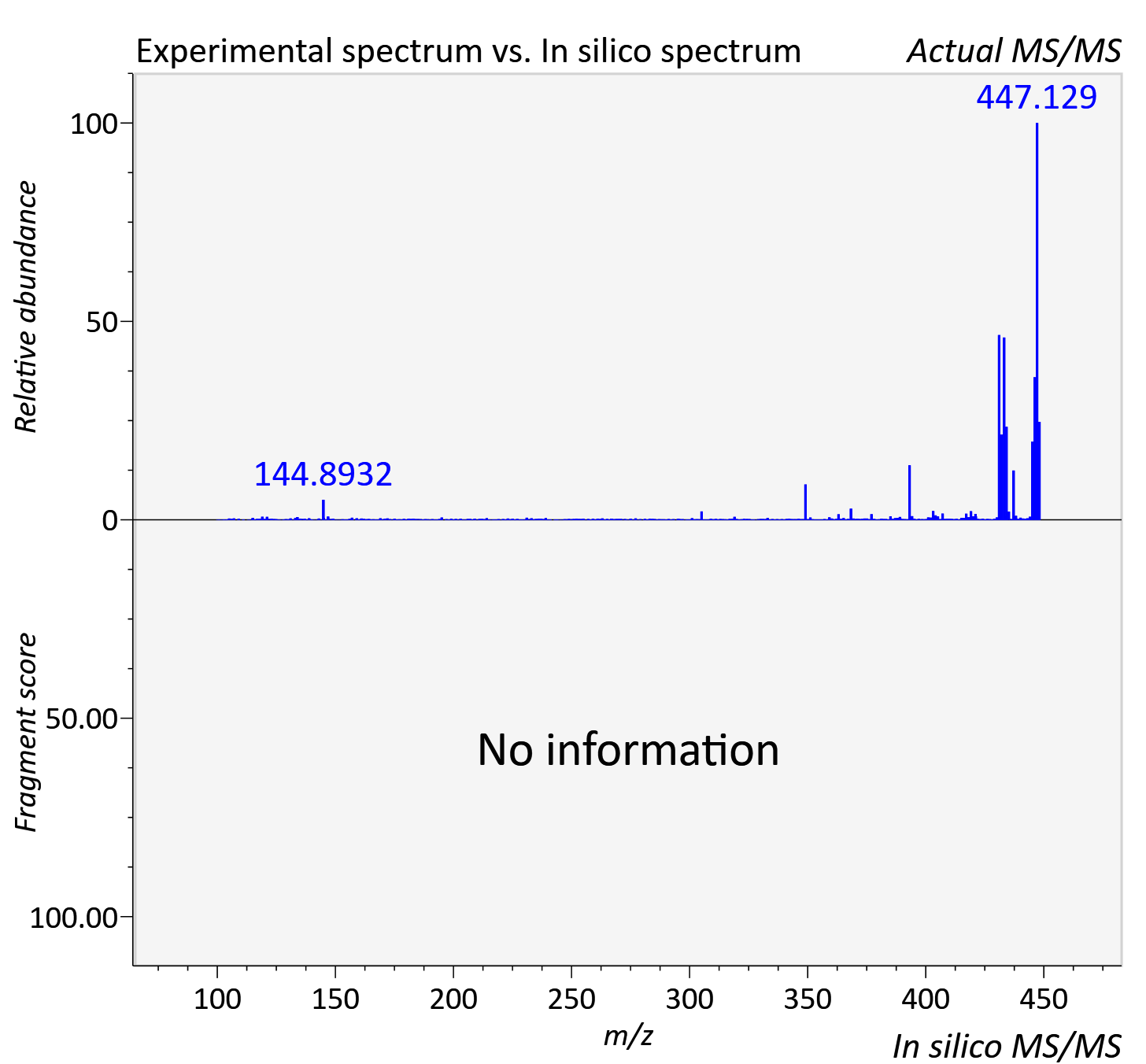
Experimental data
| Retention time | 13.15 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 448.34 | Theoretical mz | 448.342 |
| Error | 5.97 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.1661 |
Identifiers and class information
| Inchi key | FBKQAVWTYDQPAB-NUXUQDNGNA-N |
|---|---|
| Smiles | OCC1CNC2(OC3CC4C5CCC6CC(O)CCC6(C)C5CCC4(C)C3C2C)C(O)C1 |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q99720 | SIGMAR1 | Sigma opioid receptor | T46360 | SwissTargetPrediction |
| Q15125 | EBP | Anti-estrogen binding site (AEBS) | T25317 | SwissTargetPrediction and SEA |
| O43451 | MGAM | Maltase-glucoamylase | T92777 | SwissTargetPrediction |
| P10253 | GAA | Lysosomal alpha-glucosidase (by homology) | T31514 | SwissTargetPrediction |
| Q05586 | GRIN1 | Glutamate NMDA receptor; GRIN1/GRIN2B | T62276 | SEA |
| P53985 | SLC16A1 | Monocarboxylate transporter 1 (by homology) | T95842 | SwissTargetPrediction |
| P08185 | SERPINA6 | Corticosteroid binding globulin | T88452 | SEA |
| Q8TDU6 | GPBAR1 | G-protein coupled bile acid receptor 1 | T86273 | SEA |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SwissTargetPrediction |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SwissTargetPrediction |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
| O75116 | ROCK2 | Rho-associated protein kinase 2 | T06093 | SwissTargetPrediction |
| P31749 | AKT1 | Serine/threonine-protein kinase AKT | T67619 | SwissTargetPrediction |
| Q9NWZ3 | IRAK4 | Interleukin-1 receptor-associated kinase 4 | T55398 | SwissTargetPrediction |
| P31751 | AKT2 | Serine/threonine-protein kinase AKT2 | T94621 | SwissTargetPrediction |
| Q9Y243 | AKT3 | Serine/threonine-protein kinase AKT | T71266 | SwissTargetPrediction |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| P60842 | EIF4A1 | Eukaryotic initiation factor 4A-I | T86805 | SEA |
| Q9NZ70 | TAK1 | TGF-beta-activated kinase 1 | T04361 | SwissTargetPrediction |
| P51617 | IRAK1 | Interleukin-1 receptor-associated kinase 1 | T63846 | SwissTargetPrediction |
| P62508 | ESRRG | Estrogen-related receptor gamma | T59875 | SwissTargetPrediction |
| Q13224 | GRIN2B | Glutamate [NMDA] receptor subunit epsilon 2 | T76414 | SEA |
| O15399 | GRIN2D | Glutamate [NMDA] receptor subunit epsilon 4 | T56588 | SEA |
| Q14973 | SLC10A1 | Bile acid transporter | T99189 | SEA |
| P38571 | LIPA | Lysosomal acid lipase/cholesteryl ester hydrolase | T24373 | SEA |
| Q14957 | GRIN2C | Glutamate receptor ionotropic NMDA 2C | T72595 | SEA |
| O14764 | GABRD | GABA(A) receptor delta | T27376 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T46360 | DI0105 | Cough | [ICD-11: MD12] | Q99720 | SIGMAR1 |
| T25317 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q15125 | EBP |
| T92777 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | O43451 | MGAM |
| T92777 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | O43451 | MGAM |
| T31514 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P10253 | GAA |
| T62276 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q05586 | GRIN1 |
| T62276 | DI0182 | HIV-infected patients with tuberculosis | [ICD-11: 1B10-1B14] | Q05586 | GRIN1 |
| T88452 | DI0037 | Asthma | [ICD-11: CA23] | P08185 | SERPINA6 |
| T88452 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P08185 | SERPINA6 |
| T88452 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08185 | SERPINA6 |
| T86273 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | Q8TDU6 | GPBAR1 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T06093 | DI0168 | Graft-versus-host disease | [ICD-11: 4B24] | O75116 | ROCK2 |
| T67619 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P31749 | AKT1 |
| T55398 | DI0039 | Atopic eczema | [ICD-11: EA80] | Q9NWZ3 | IRAK4 |
| T55398 | DI0109 | Cutaneous lupus erythematosus | [ICD-11: EB50-EB5Z] | Q9NWZ3 | IRAK4 |
| T55398 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q9NWZ3 | IRAK4 |
| T55398 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | Q9NWZ3 | IRAK4 |
| T55398 | DI0351 | Psoriasis | [ICD-11: EA90] | Q9NWZ3 | IRAK4 |
| T55398 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9NWZ3 | IRAK4 |
| T71266 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q9Y243 | AKT3 |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T86805 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P60842 | EIF4A1 |
| T04361 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | Q9NZ70 | TAK1 |
| T76414 | DI0024 | Alpha-1-antitrypsin deficiency | [ICD-11: 5C5A] | Q13224 | GRIN2B |
| T56588 | DI0296 | Neurodegenerative disorder | [ICD-11: 8A20-8A23] | O15399 | GRIN2D |
| T99189 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | Q14973 | SLC10A1 |
| T24373 | DI0132 | Enzyme deficiency | [ICD-11: 5C51-5C57] | P38571 | LIPA |
| T72595 | DI0296 | Neurodegenerative disorder | [ICD-11: 8A20-8A23] | Q14957 | GRIN2C |
| T27376 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O14764 | GABRD |
| T27376 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | O14764 | GABRD |
| T27376 | DI0256 | Mental/behavioural/neurodevelopmental disorder | [ICD-11: 6E20-6E8Z] | O14764 | GABRD |
