Compound details
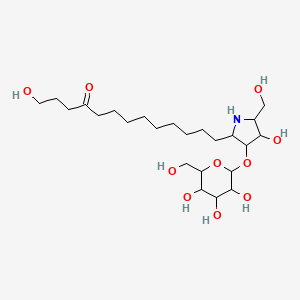
Broussonetine A
| Compound ID | CDAMM02682 |
|---|---|
| Common name | Broussonetine A | IUPAC name | 1-hydroxy-13-[4-hydroxy-5-(hydroxymethyl)-3-[3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxypyrrolidin-2-yl]tridecan-4-one |
| Molecular formula | C24H45NO10 |
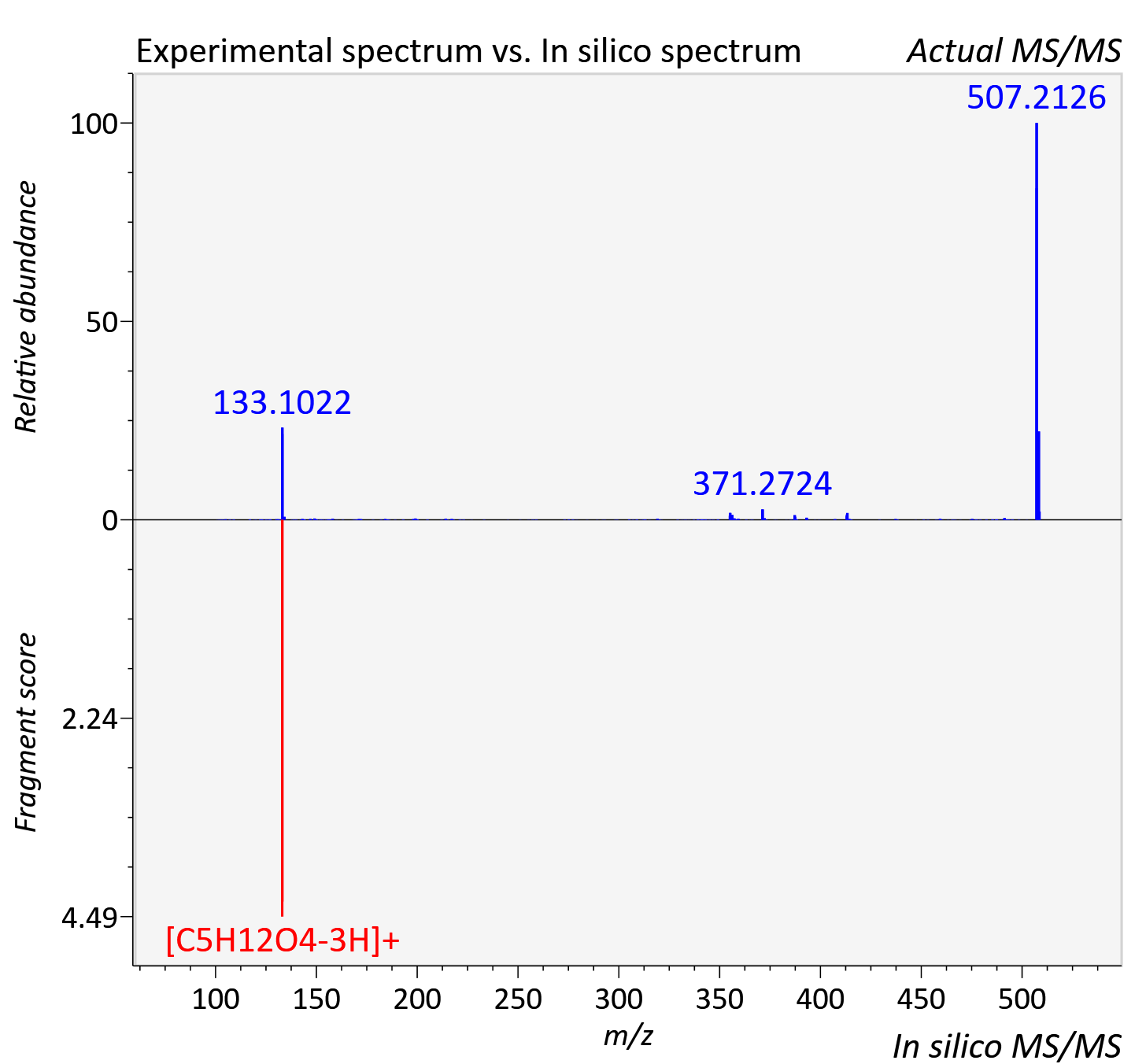
Experimental data
| Retention time | 11.97 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 508.309 | Theoretical mz | 508.311 |
| Error | 3.69 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.0407 |
Identifiers and class information
| Inchi key | LYIJINGICVFWEP-OWXBEFEHNA-N |
|---|---|
| Smiles | O=C(CCCO)CCCCCCCCCC1NC(CO)C(O)C1OC2OC(CO)C(O)C(O)C2O |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P06280 | GLA | Alpha-galactosidase A | T61339 | SEA |
| P06280 | GLA | Alpha-galactosidase A | T61339 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| Q9BZP6 | CHIA | Acidic mammalian chitinase (by homology) | T51597 | SEA |
| P32320 | CDA | Cytidine deaminase (by homology) | T79027 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| P05230 | FGF1 | Acidic fibroblast growth factor | T18639 | SEA |
| P09038 | FGF2 | Basic fibroblast growth factor | T31621 | SEA |
| P16109 | SELP | P-selectin | T10965 | SEA |
| P14151 | SELL | Leukocyte adhesion molecule-1 | T60526 | SEA |
| P17931 | LGALS3 | Galectin-3 | T72038 | SEA |
| P04746 | AMY2A | Pancreatic alpha-amylase | T86918 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| O00142 | TK2 | Thymidine kinase, mitochondrial | T22976 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T79027 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P32320 | CDA |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T18639 | DI0081 | Chronic arterial occlusive disease | [ICD-11: BD4Z] | P05230 | FGF1 |
| T18639 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P05230 | FGF1 |
| T31621 | DI0005 | Acne vulgaris | [ICD-11: ED80] | P09038 | FGF2 |
| T10965 | DI0088 | Circulatory system disease | [ICD-11: BE2Z] | P16109 | SELP |
| T10965 | DI0381 | Sickle-cell disorder | [ICD-11: 3A51] | P16109 | SELP |
| T60526 | DI0037 | Asthma | [ICD-11: CA23] | P14151 | SELL |
| T60526 | DI0088 | Circulatory system disease | [ICD-11: BE2Z] | P14151 | SELL |
| T72038 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P17931 | LGALS3 |
| T72038 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | P17931 | LGALS3 |
| T72038 | DI0252 | Melanoma | [ICD-11: 2C30] | P17931 | LGALS3 |
| T72038 | DI0302 | Non-alcoholic fatty liver disease | [ICD-11: DB92] | P17931 | LGALS3 |
| T72038 | DI0351 | Psoriasis | [ICD-11: EA90] | P17931 | LGALS3 |
| T86918 | DI0110 | Cystic fibrosis | [ICD-11: CA25] | P04746 | AMY2A |
| T86918 | DI0328 | Pancreatic malfunction | [ICD-11: DC30-DC3Z] | P04746 | AMY2A |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T22976 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | O00142 | TK2 |
