Compound details
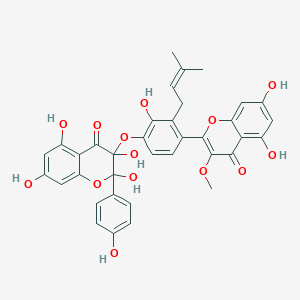
Podoverine C
| Compound ID | CDAMM02625 |
|---|---|
| Common name | Podoverine C | IUPAC name | 3-[4-(5,7-dihydroxy-3-methoxy-4-oxochromen-2-yl)-2-hydroxy-3-(3-methylbut-2-enyl)phenoxy]-2,3,5,7-tetrahydroxy-2-(4-hydroxyphenyl)chromen-4-one |
| Molecular formula | C36H30O14 |
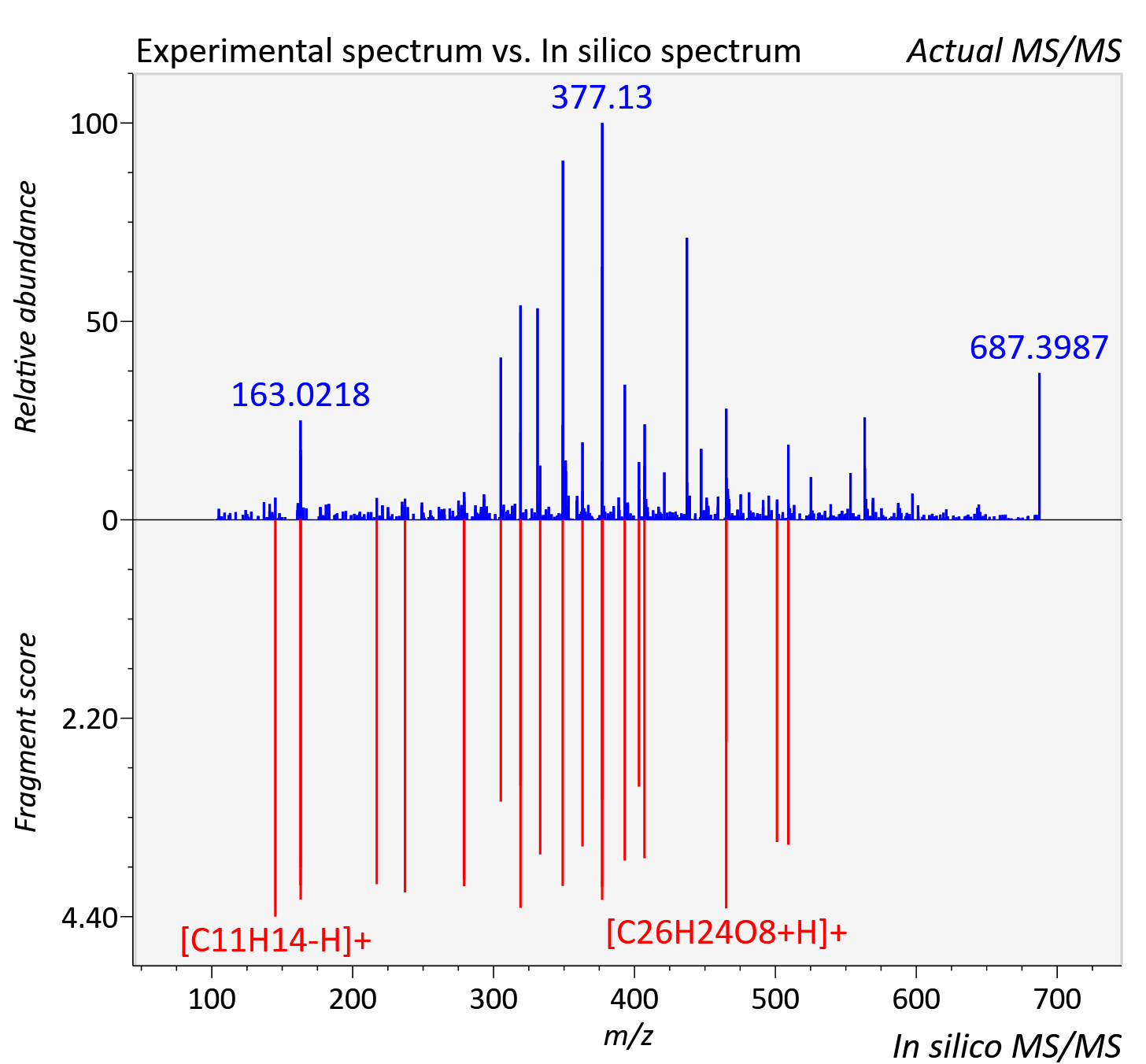
Experimental data
| Retention time | 7.63 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 687.179 | Theoretical mz | 687.171 |
| Error | 11.39 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.0597 |
Identifiers and class information
| Inchi key | JTYQTVPZTBVCSU-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1C(OC)=C(OC=2C=C(O)C=C(O)C12)C=3C=CC(OC4(O)C(=O)C=5C(O)=CC(O)=CC5OC4(O)C6=CC=C(O)C=C6)=C(O)C3CC=C(C)C |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P19838 | NFKB1 | Nuclear factor NF-kappa-B p105 subunit | T83145 | SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| P33527 | ABCC1 | Multidrug resistance-associated protein 1 | T11288 | SEA |
| Q9UNQ0 | ABCG2 | ATP-binding cassette sub-family G member 2 | T56556 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| Q9NPH5 | NOX4 | NADPH oxidase 4 | T29741 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
| P47989 | XDH | Xanthine dehydrogenase | T40954 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| Q04760 | GLO1 | Glyoxalase I | T88285 | SEA |
| Q9HC98 | NEK6 | Serine/threonine-protein kinase NEK6 | T78992 | SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T11288 | DI0167 | Gout | [ICD-11: FA25] | P33527 | ABCC1 |
| T56556 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9UNQ0 | ABCG2 |
| T56556 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9UNQ0 | ABCG2 |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T29741 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9NPH5 | NOX4 |
| T40954 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | P47989 | XDH |
| T40954 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P47989 | XDH |
| T88285 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | Q04760 | GLO1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
