Compound details
Jurubine
| Compound ID | CDAMM02621 |
|---|---|
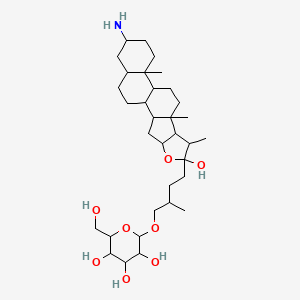
| Common name | Jurubine | IUPAC name | 2-[4-(16-amino-6-hydroxy-7,9,13-trimethyl-5-oxapentacyclo[10.8.0.02,9.04,8.013,18]icosan-6-yl)-2-methylbutoxy]-6-(hydroxymethyl)oxane-3,4,5-triol |
| Molecular formula | C33H57NO8 |
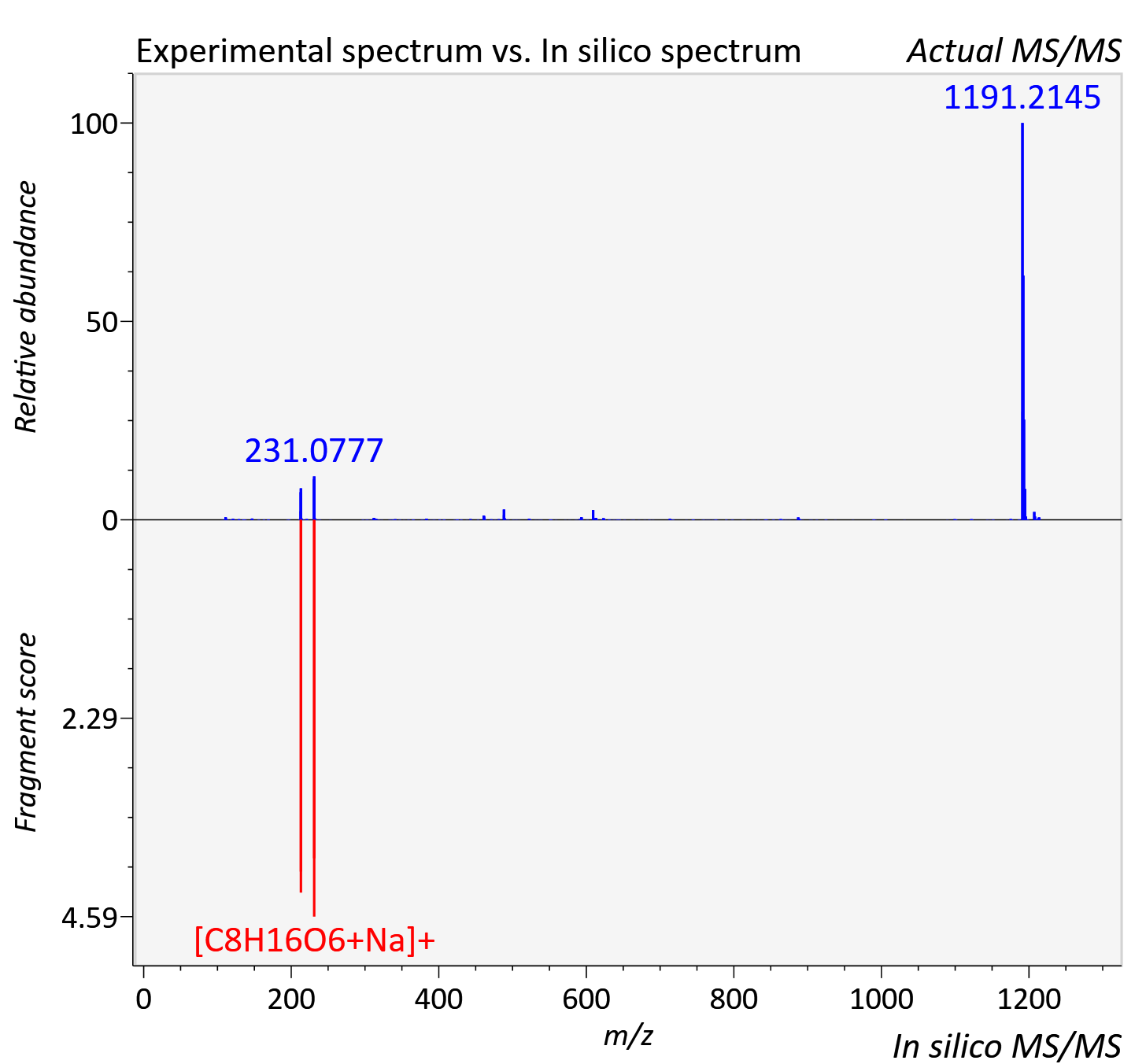
Experimental data
| Retention time | 14.84 |
|---|---|
| Adduct | [2M+Na]+ |
| Actual mz | 1213.8 | Theoretical mz | 1213.81 |
| Error | 1.83 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.3135 |
Identifiers and class information
| Inchi key | YEWUMIMAJWFDQG-UHFFFAOYNA-N |
|---|---|
| Smiles | OCC1OC(OCC(C)CCC2(O)OC3CC4C5CCC6CC(N)CCC6(C)C5CCC4(C)C3C2C)C(O)C(O)C1O |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P08185 | SERPINA6 | Corticosteroid binding globulin | T88452 | SEA |
| Q8TDU6 | GPBAR1 | G-protein coupled bile acid receptor 1 | T86273 | SEA |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
| P05230 | FGF1 | Acidic fibroblast growth factor | T18639 | SEA |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| P60842 | EIF4A1 | Eukaryotic initiation factor 4A-I | T86805 | SEA |
| O15439 | ABCC4 | Multidrug resistance protein 4 | T39919 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T88452 | DI0037 | Asthma | [ICD-11: CA23] | P08185 | SERPINA6 |
| T88452 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P08185 | SERPINA6 |
| T88452 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08185 | SERPINA6 |
| T86273 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | Q8TDU6 | GPBAR1 |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T18639 | DI0081 | Chronic arterial occlusive disease | [ICD-11: BD4Z] | P05230 | FGF1 |
| T18639 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P05230 | FGF1 |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T86805 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P60842 | EIF4A1 |
