Compound details
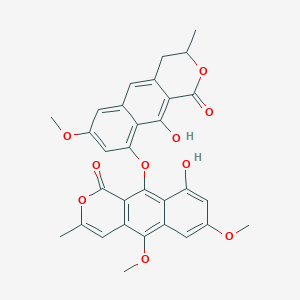
Planifolin
| Compound ID | CDAMM02615 |
|---|---|
| Common name | Planifolin | IUPAC name | 9-hydroxy-10-[(10-hydroxy-7-methoxy-3-methyl-1-oxo-3,4-dihydrobenzo[g]isochromen-9-yl)oxy]-5,7-dimethoxy-3-methylbenzo[g]isochromen-1-one |
| Molecular formula | C31H26O10 |
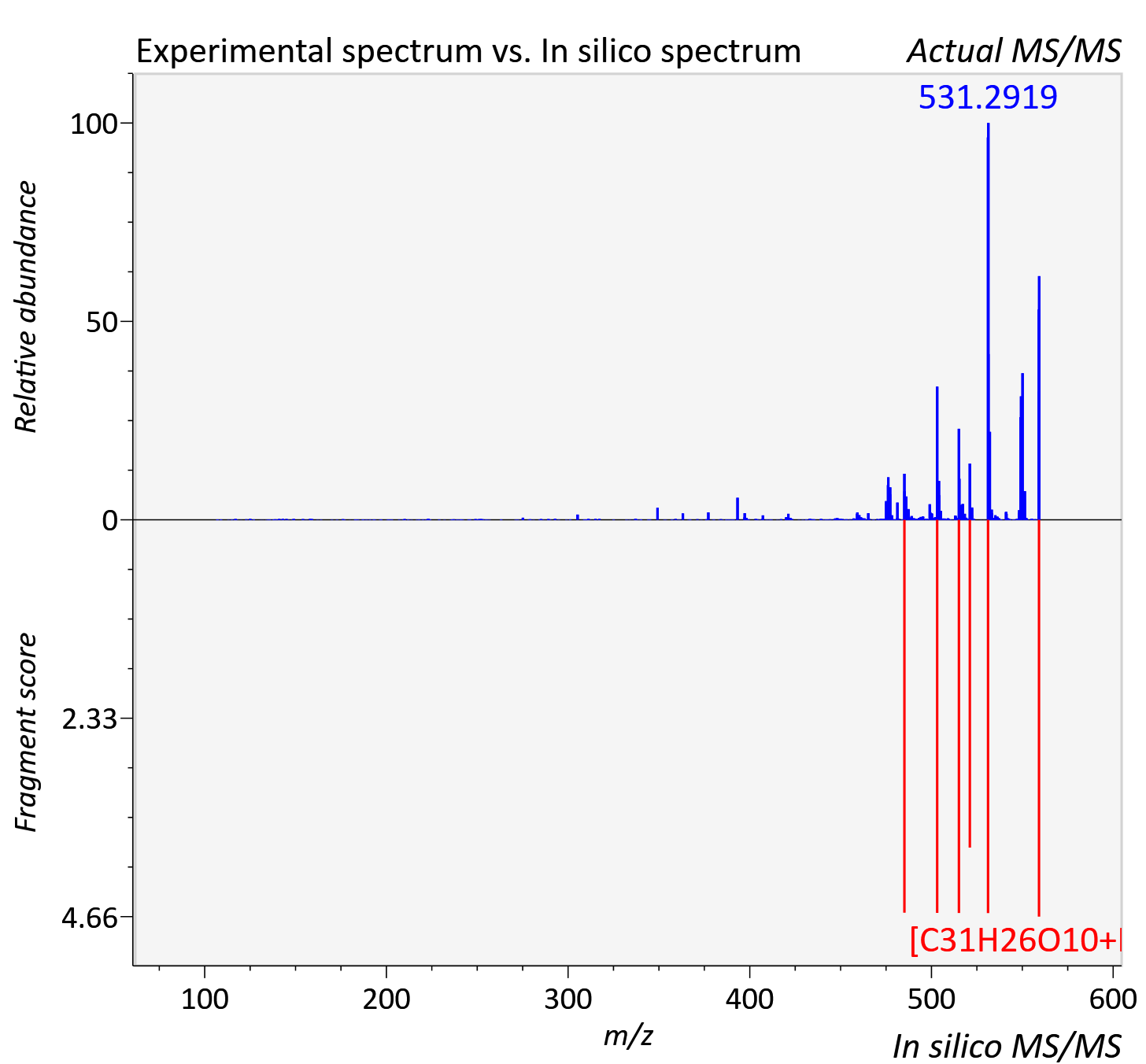
Experimental data
| Retention time | 12.54 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 559.161 | Theoretical mz | 559.16 |
| Error | 1.83 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.7337 |
Identifiers and class information
| Inchi key | AFCSPOKMAFUNHQ-UGPWUYPHNA-N |
|---|---|
| Smiles | O=C1OC(=CC=2C(OC)=C3C=C(OC)C=C(O)C3=C(OC4=CC(OC)=CC=5C=C6C(C(=O)OC(C)C6)=C(O)C45)C12)C |
| Superclass | Organoheterocyclic compounds |
| Class | Naphthopyrans |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P37023 | ACVRL1 | Serine/threonine-protein kinase receptor R3 | T36959 | SEA |
| P16234 | PDGFRA | Platelet-derived growth factor receptor | T53524 | SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SEA |
| Q9NZ70 | TAK1 | TGF-beta-activated kinase 1 | T04361 | SEA |
| Q9UK17 | KCND3 | Voltage-gated potassium channel Kv4.3 | T74500 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T36959 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P37023 | ACVRL1 |
| T53524 | DI0160 | Gastrointestinal stromal tumour | [ICD-11: 2B5B] | P16234 | PDGFRA |
| T53524 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P16234 | PDGFRA |
| T53524 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P16234 | PDGFRA |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T04361 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | Q9NZ70 | TAK1 |
| T74500 | DI0004 | Acidosis | [ICD-11: 5C73] | Q9UK17 | KCND3 |
