Compound details
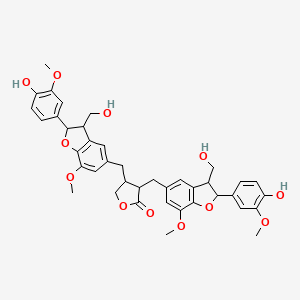
Lappaol F
| Compound ID | CDAMM02614 |
|---|---|
| Common name | Lappaol F | IUPAC name | 3,4-bis[[2-(4-hydroxy-3-methoxyphenyl)-3-(hydroxymethyl)-7-methoxy-2,3-dihydro-1-benzofuran-5-yl]methyl]oxolan-2-one |
| Molecular formula | C40H42O12 |
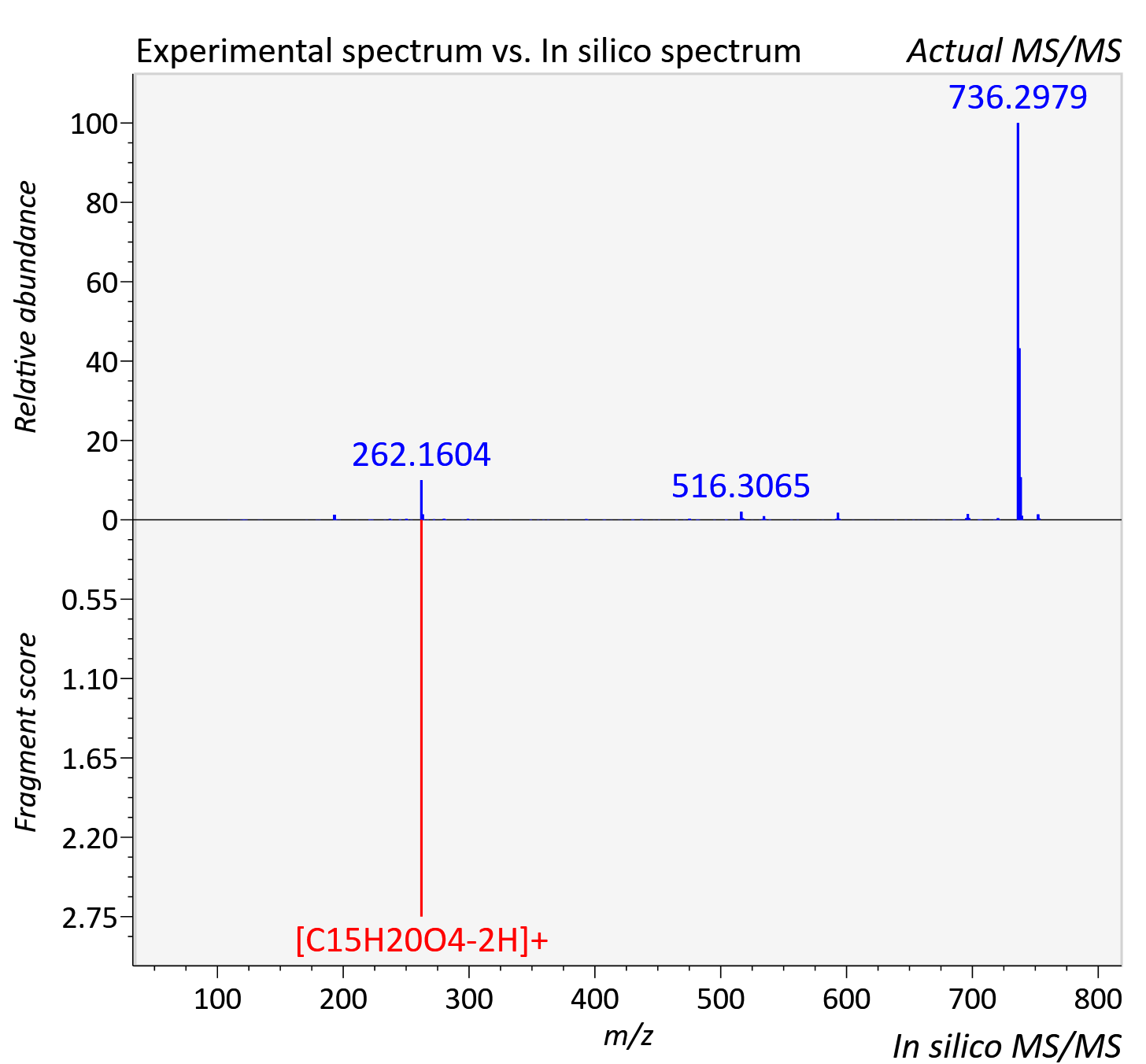
Experimental data
| Retention time | 16.21 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 753.232 | Theoretical mz | 753.231 |
| Error | 1.78 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.0326 |
Identifiers and class information
| Inchi key | YXNKOCZXAVTXTG-OXHLKWRSNA-N |
|---|---|
| Smiles | O=C1OCC(CC2=CC(OC)=C3OC(C4=CC=C(O)C(OC)=C4)C(C3=C2)CO)C1CC5=CC(OC)=C6OC(C7=CC=C(O)C(OC)=C7)C(C6=C5)CO |
| Superclass | Phenylpropanoids and polyketides |
| Class | 2-arylbenzofuran flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| P51843 | NR0B1 | Orphan nuclear receptor DAX-1 | T23191 | SEA |
| Q8N183 | NDUFAF2 | HUMAN NADH:ubiquinone oxidoreductase complex assembly factor 2 | T01453 | SEA |
| Q9P0J0 | NDUFA13 | Mitochondrial complex I | T82391 | SEA |
| Q9Y6M9 | NDUFB9 | HUMAN NADH:ubiquinone oxidoreductase subunit B9 | T01465 | SEA |
| Q8N183 | NDUFAF2 | HUMAN NADH:ubiquinone oxidoreductase complex assembly factor 2 | T01453 | SEA |
| P03897 | MT-ND3 | NADH dehydrogenase | T77195 | SEA |
