Compound details
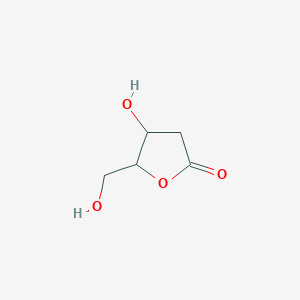
2-Deoxy-L-ribono-1,4-lactone
| Compound ID | CDAMM02591 |
|---|---|
| Common name | 2-Deoxy-L-ribono-1,4-lactone | IUPAC name | 4-hydroxy-5-(hydroxymethyl)oxolan-2-one |
| Molecular formula | C5H8O4 |
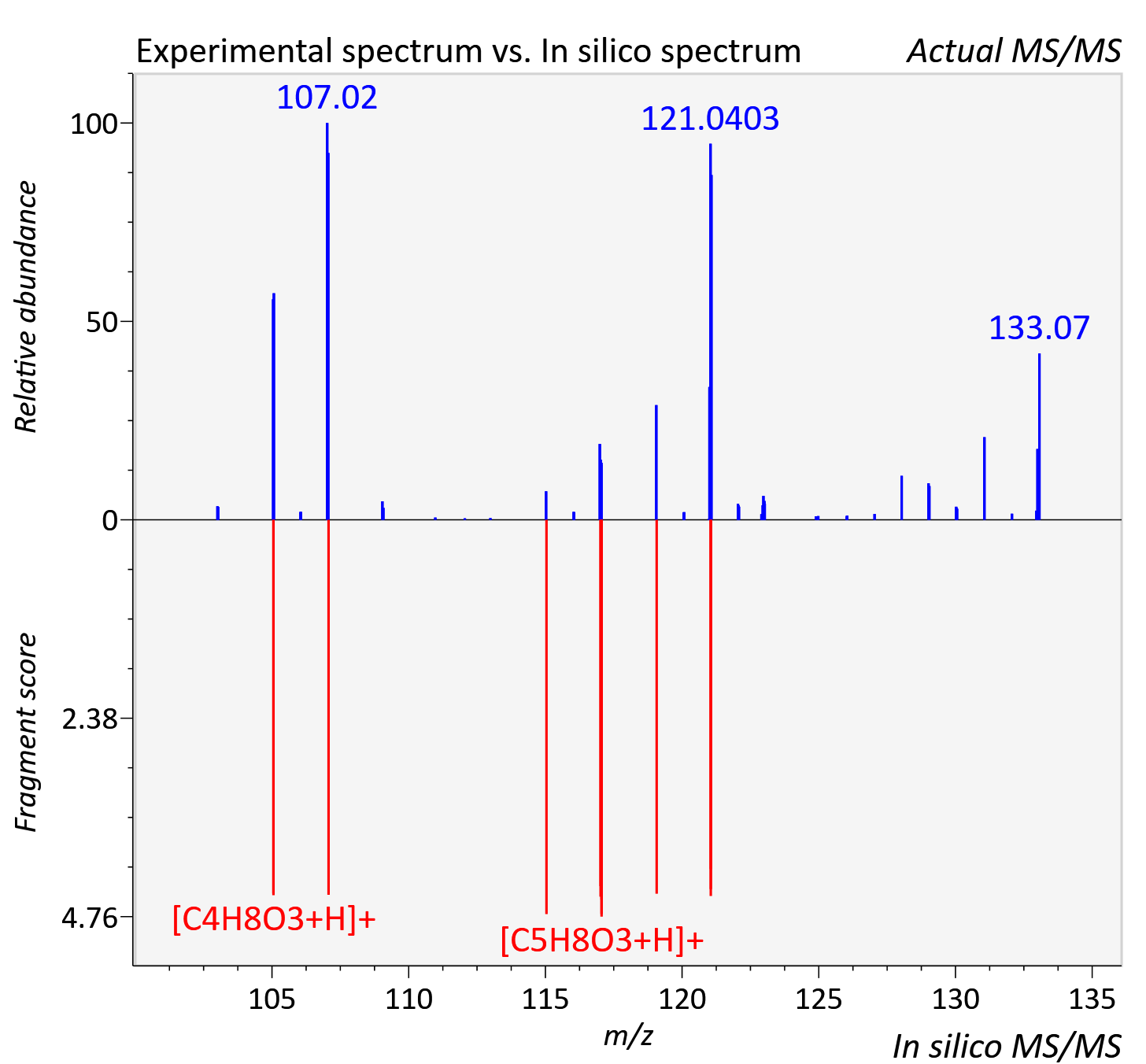
Experimental data
| Retention time | 9.54 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 133.051 | Theoretical mz | 133.049 |
| Error | 11.45 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.7052 |
Identifiers and class information
| Inchi key | YIXDEYPPAGPYDP-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC(CO)C(O)C1 |
| Superclass | Organoheterocyclic compounds |
| Class | Lactones |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T30081 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P04183 | TK1 |
| T30081 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | P04183 | TK1 |
| T79027 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P32320 | CDA |
