Compound details
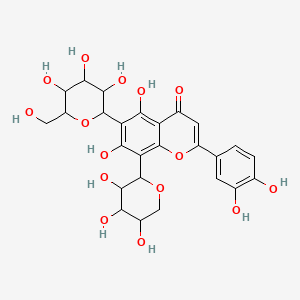
Neocarlinoside
| Compound ID | CDAMM02579 |
|---|---|
| Common name | Neocarlinoside | IUPAC name | 2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-6-[3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]-8-(3,4,5-trihydroxyoxan-2-yl)chromen-4-one |
| Molecular formula | C26H28O15 |
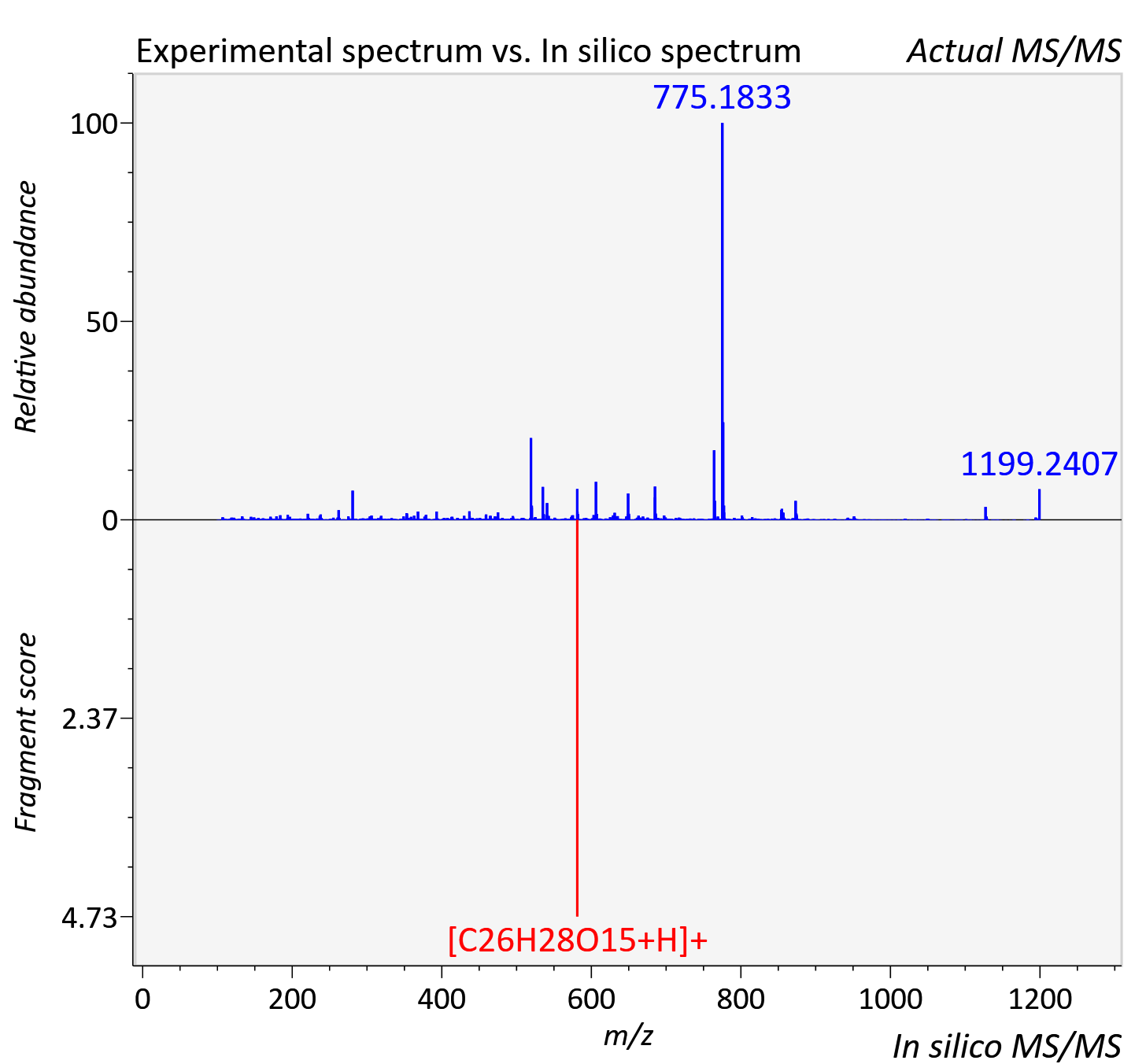
Experimental data
| Retention time | 17.37 |
|---|---|
| Adduct | [2M+K]+ |
| Actual mz | 1199.24 | Theoretical mz | 1199.25 |
| Error | 10.97 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.6551 |
Identifiers and class information
| Inchi key | XBGYTZHKGMCEGE-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1C=C(OC2=C1C(O)=C(C(O)=C2C3OCC(O)C(O)C3O)C4OC(CO)C(O)C(O)C4O)C=5C=CC(O)=C(O)C5 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Flavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P15121 | AKR1B1 | Aldose reductase | T26623 | SwissTargetPrediction and SEA |
| O14746 | TERT | Telomerase reverse transcriptase | T86052 | SEA |
| P15692 | VEGFA | Vascular endothelial growth factor A | T20761 | SEA |
| Q07002 | CDK18 | Serine/threonine-protein kinase PCTAIRE-3 | T27577 | SEA |
| P01112 | HRAS | Transforming protein p21/H-Ras-1 | T71081 | SEA |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| P49763 | PGF | Placenta growth factor | T70792 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| Q9NPH5 | NOX4 | NADPH oxidase 4 | T29741 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
| P47989 | XDH | Xanthine dehydrogenase | T40954 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| Q04760 | GLO1 | Glyoxalase I | T88285 | SEA |
| P20248 | CCNA2 | CDK2/Cyclin A | T58470 | SEA |
| Q14004 | CDK13 | Cyclin-dependent kinase 13 | T74839 | SEA |
| Q16512 | PKN1 | Protein kinase N1 | T60003 | SEA |
| Q9HC98 | NEK6 | Serine/threonine-protein kinase NEK6 | T78992 | SEA |
| P09382 | LGALS1 | Galectin-1 | T09544 | SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| Q9NZ08 | ERAP1 | Endoplasmic reticulum aminopeptidase 1 | T72849 | SEA |
| P05231 | IL6 | Interleukin-6 | T32578 | SEA |
| P21860 | ERBB3 | Receptor tyrosine-protein kinase erbB-3 | T86350 | SEA |
| Q13705 | ACVR2B | Activin receptor type-2B | T80338 | SEA |
| Q99417 | MYCBP | MYCBP messenger RNA | T37298 | SEA |
| P16220 | CREB1 | Cyclic AMP-responsive element-binding protein | T92098 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
| P30837 | ALDH1B1 | Acetaldehyde dehydrogenase | T99641 | SEA |
| P27037 | ACVR2A | Activin receptor type IIA | T47366 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T26623 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P15121 | AKR1B1 |
| T26623 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P15121 | AKR1B1 |
| T86052 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | O14746 | TERT |
| T86052 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | O14746 | TERT |
| T86052 | DI0326 | Pancreatic cancer | [ICD-11: 2C10] | O14746 | TERT |
| T20761 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P15692 | VEGFA |
| T20761 | DI0365 | Retinopathy | [ICD-11: 9B71] | P15692 | VEGFA |
| T20761 | DI0430 | Vascular system developmental anomaly | [ICD-11: LA90] | P15692 | VEGFA |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T70792 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P49763 | PGF |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T29741 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9NPH5 | NOX4 |
| T40954 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | P47989 | XDH |
| T40954 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P47989 | XDH |
| T88285 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | Q04760 | GLO1 |
| T58470 | DI0363 | Retina cancer | [ICD-11: 2D02] | P20248 | CCNA2 |
| T74839 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q14004 | CDK13 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T32578 | DI0028 | Anemia | [ICD-11: 3A00-3A9Z] | P05231 | IL6 |
| T86350 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P21860 | ERBB3 |
| T86350 | DI0095 | Colorectal cancer | [ICD-11: 2B91] | P21860 | ERBB3 |
| T86350 | DI0172 | Head and neck cancer | [ICD-11: 2D42] | P21860 | ERBB3 |
| T86350 | DI0238 | Lung cancer | [ICD-11: 2C25] | P21860 | ERBB3 |
| T86350 | DI0246 | Malignant mesenchymal neoplasm | [ICD-11: 2B5D-2B5Y] | P21860 | ERBB3 |
| T86350 | DI0259 | Metastatic lymph node neoplasm | [ICD-11: 2D60] | P21860 | ERBB3 |
| T86350 | DI0326 | Pancreatic cancer | [ICD-11: 2C10] | P21860 | ERBB3 |
| T86350 | DI0351 | Psoriasis | [ICD-11: EA90] | P21860 | ERBB3 |
| T86350 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P21860 | ERBB3 |
| T86350 | DI0393 | Squamous cell carcinoma | [ICD-11: 2B60-2D01] | P21860 | ERBB3 |
| T80338 | DI0295 | Nervous system paraneoplastic/autoimmune disorder | [ICD-11: 8E4A] | Q13705 | ACVR2B |
| T37298 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q99417 | MYCBP |
| T99641 | DI0396 | Substance abuse | [ICD-11: 6C40] | P30837 | ALDH1B1 |
| T47366 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P27037 | ACVR2A |
| T47366 | DI0198 | Idiopathic inflammatory myopathy | [ICD-11: 4A41] | P27037 | ACVR2A |
