Compound details
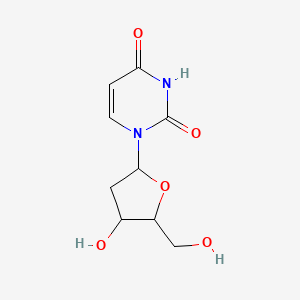
Deoxyuridine
| Compound ID | CDAMM02512 |
|---|---|
| Common name | Deoxyuridine | IUPAC name | 1-[4-hydroxy-5-(hydroxymethyl)oxolan-2-yl]pyrimidine-2,4-dione |
| Molecular formula | C9H12N2O5 |
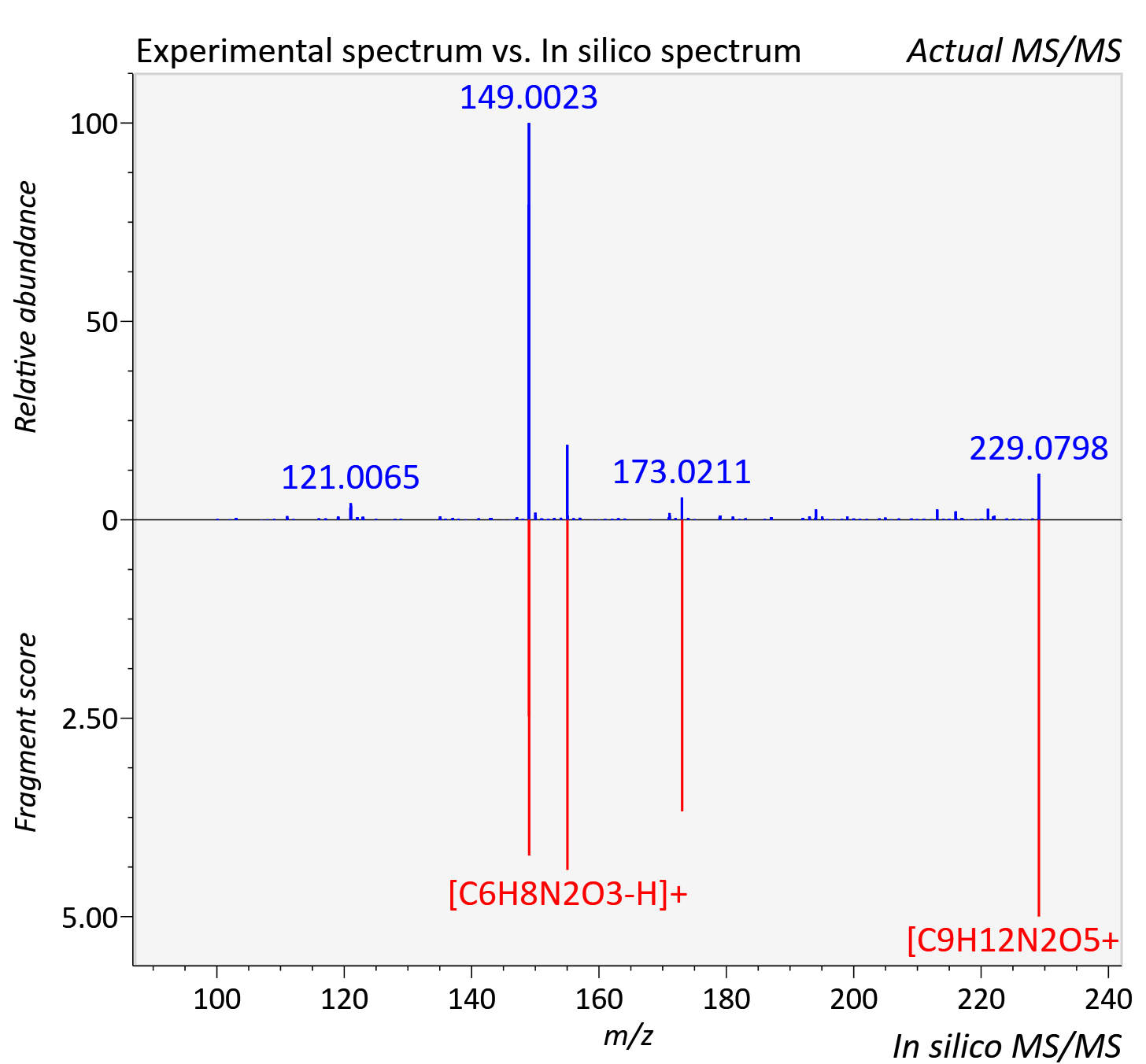
Experimental data
| Retention time | 10.71 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 229.081 | Theoretical mz | 229.082 |
| Error | 3.39 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.5872 |
Identifiers and class information
| Inchi key | MXHRCPNRJAMMIM-SHYZEUOFSA-N |
|---|---|
| Smiles | O=C1C=CN(C(=O)N1)C2OC(CO)C(O)C2 |
| Superclass | Nucleosides, nucleotides, and analogues |
| Class | Pyrimidine nucleosides |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P04818 | TYMS | Thymidylate synthase (by homology) | T98397 | SEA |
| P04183 | TK1 | Thymidine kinase, cytosolic | T30081 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| P32320 | CDA | Cytidine deaminase (by homology) | T79027 | SEA |
| P19971 | TYMP | Thymidine phosphorylase | T59929 | SEA |
| P41231 | P2RY2 | Purinergic receptor P2Y2 | T93515 | SEA |
| Q15077 | P2RY6 | Pyrimidinergic receptor P2Y6 | T90782 | SEA |
| P23919 | DTYMK | Thymidylate kinase | T94400 | SEA |
| O00142 | TK2 | Thymidine kinase, mitochondrial | T22976 | SEA |
| Q15391 | P2RY14 | Purinergic receptor P2Y14 | T16555 | SEA |
| P28340 | POLD1 | DNA polymerase delta subunit 1 | T02258 | SEA |
| Q96SW2 | CRBN | Protein cereblon | T27861 | SEA |
| Q13304 | GPR17 | Uracil nucleotide/cysteinyl leukotriene receptor | T33902 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T98397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P04818 | TYMS |
| T98397 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P04818 | TYMS |
| T30081 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P04183 | TK1 |
| T30081 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | P04183 | TK1 |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T79027 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P32320 | CDA |
| T59929 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P19971 | TYMP |
| T93515 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P41231 | P2RY2 |
| T90782 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q15077 | P2RY6 |
| T22976 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | O00142 | TK2 |
| T02258 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P28340 | POLD1 |
| T27861 | DI0060 | Brain cancer | [ICD-11: 2A00] | Q96SW2 | CRBN |
| T27861 | DI0123 | Diffuse large B-cell lymphoma | [ICD-11: 2A81] | Q96SW2 | CRBN |
| T27861 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q96SW2 | CRBN |
| T27861 | DI0239 | Lupus erythematosus | [ICD-11: 4A40] | Q96SW2 | CRBN |
| T27861 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q96SW2 | CRBN |
| T27861 | DI0249 | Mature B-cell leukaemia | [ICD-11: 2A82] | Q96SW2 | CRBN |
| T27861 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | Q96SW2 | CRBN |
| T27861 | DI0368 | Sarcoidosis | [ICD-11: 4B20] | Q96SW2 | CRBN |
| T27861 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q96SW2 | CRBN |
