Compound details
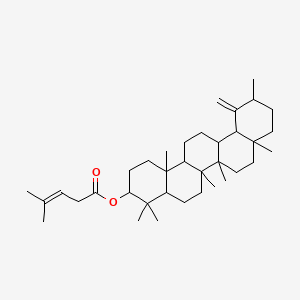
pergularine A
| Compound ID | CDAMM02499 |
|---|---|
| Common name | pergularine A | IUPAC name | (4,4,6a,6b,8a,11,14b-heptamethyl-12-methylidene-1,2,3,4a,5,6,6a,7,8,9,10,11,12a,13,14,14a-hexadecahydropicen-3-yl) 4-methylpent-3-enoate |
| Molecular formula | C36H58O2 |
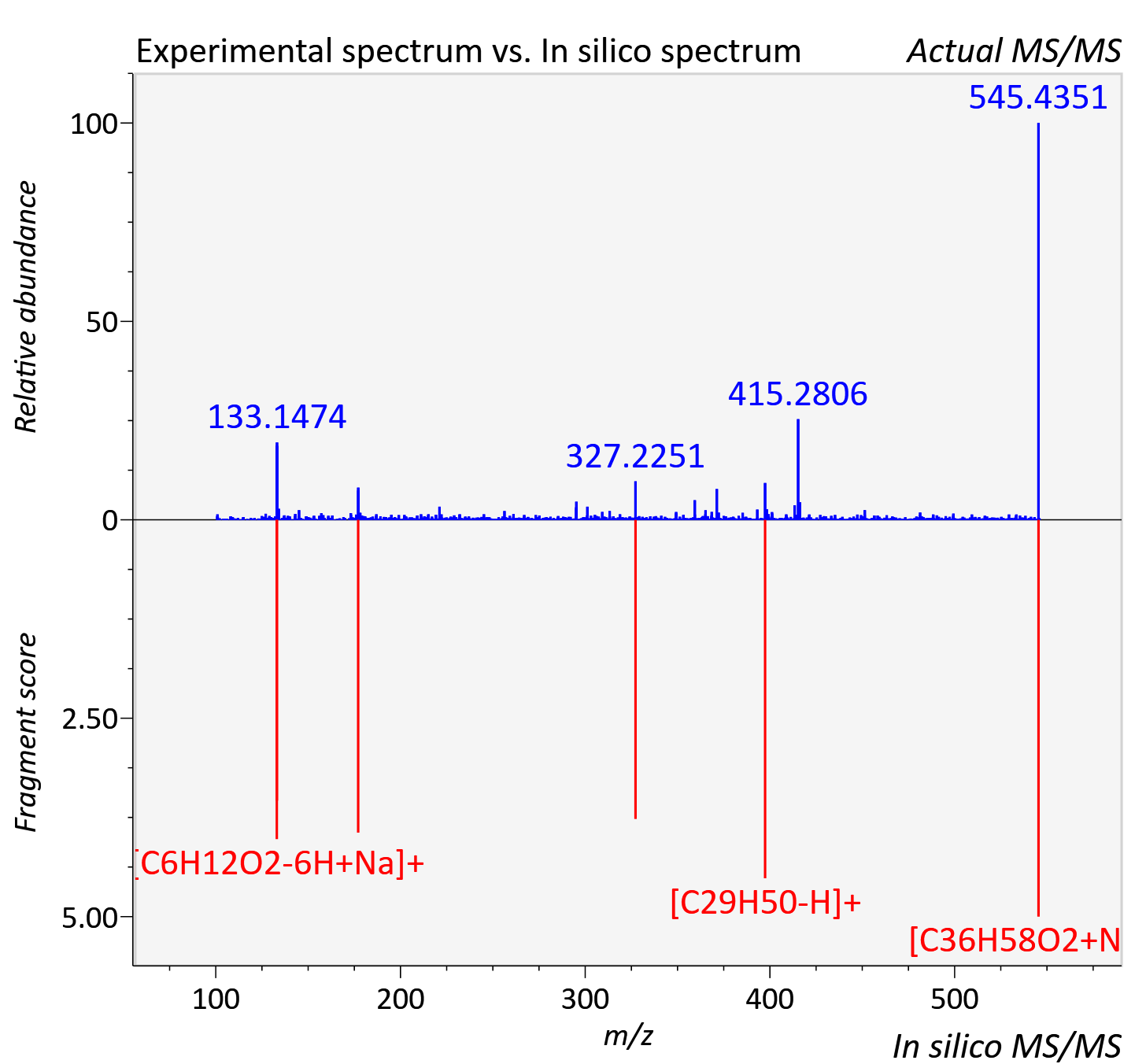
Experimental data
| Retention time | 12.59 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 545.433 | Theoretical mz | 545.433 |
| Error | 0.52 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.4457 |
Identifiers and class information
| Inchi key | SOKIHCAMPJLTAJ-XWFTXFNQSA-N |
|---|---|
| Smiles | O=C(OC1CCC2(C)C(CCC3(C)C2CCC4C5C(=C)C(C)CCC5(C)CCC43C)C1(C)C)CC=C(C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SwissTargetPrediction |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction and SEA |
| P17706 | PTPN2 | T-cell protein-tyrosine phosphatase | T49156 | SEA |
| P28845 | HSD11B1 | 11-beta-hydroxysteroid dehydrogenase 1 | T65200 | SwissTargetPrediction |
| Q8TDU6 | GPBAR1 | G-protein coupled bile acid receptor 1 | T86273 | SEA |
| Q9Y233 | PDE10A | Phosphodiesterase 10A (by homology) | T84133 | SwissTargetPrediction |
| P30518 | AVPR2 | Vasopressin V2 receptor (by homology) | T66237 | SwissTargetPrediction |
| P05093 | CYP17A1 | Cytochrome P450 17A1 | T89041 | SwissTargetPrediction |
| O43614 | HCRTR2 | Orexin receptor 2 | T69485 | SwissTargetPrediction |
| O43613 | HCRTR1 | Orexin receptor 1 | T73482 | SwissTargetPrediction |
| P25116 | F2R | Proteinase-activated receptor 1 | T36483 | SwissTargetPrediction |
| P01584 | IL1B | Interleukin-1 beta | T42000 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T49156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P17706 | PTPN2 |
| T65200 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | P28845 | HSD11B1 |
| T65200 | DI0239 | Lupus erythematosus | [ICD-11: 4A40] | P28845 | HSD11B1 |
| T65200 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P28845 | HSD11B1 |
| T86273 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | Q8TDU6 | GPBAR1 |
| T84133 | DI0079 | Choreiform disorder | [ICD-11: 8A01] | Q9Y233 | PDE10A |
| T84133 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q9Y233 | PDE10A |
| T84133 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | Q9Y233 | PDE10A |
| T84133 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q9Y233 | PDE10A |
| T84133 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9Y233 | PDE10A |
| T66237 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P30518 | AVPR2 |
| T66237 | DI0041 | Autism spectrum disorder | [ICD-11: 6A02] | P30518 | AVPR2 |
| T66237 | DI0131 | Enuresis | [ICD-11: 6C00] | P30518 | AVPR2 |
| T66237 | DI0194 | Hypo-osmolality/hyponatraemia | [ICD-11: 5C72] | P30518 | AVPR2 |
| T66237 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P30518 | AVPR2 |
| T66237 | DI0338 | Polyuria | [ICD-11: MF55] | P30518 | AVPR2 |
| T89041 | DI0346 | Prostate cancer | [ICD-11: 2C82] | P05093 | CYP17A1 |
| T69485 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | O43614 | HCRTR2 |
| T73482 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O43613 | HCRTR1 |
| T73482 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | O43613 | HCRTR1 |
| T73482 | DI0317 | Opioid use disorder | [ICD-11: 6C43] | O43613 | HCRTR1 |
| T36483 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P25116 | F2R |
| T42000 | DI0267 | Mineral excesses | [ICD-11: 5B91] | P01584 | IL1B |
| T42000 | DI0269 | Monogenic autoinflammatory syndrome | [ICD-11: 4A60] | P01584 | IL1B |
| T42000 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P01584 | IL1B |
| T42000 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P01584 | IL1B |
