Compound details
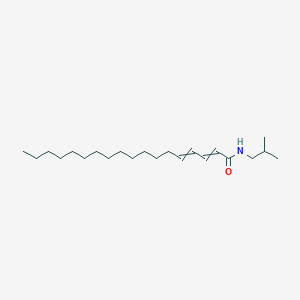
Pipericine
| Compound ID | CDAMM02433 |
|---|---|
| Common name | Pipericine | IUPAC name | N-(2-methylpropyl)octadeca-2,4-dienamide |
| Molecular formula | C22H41NO |
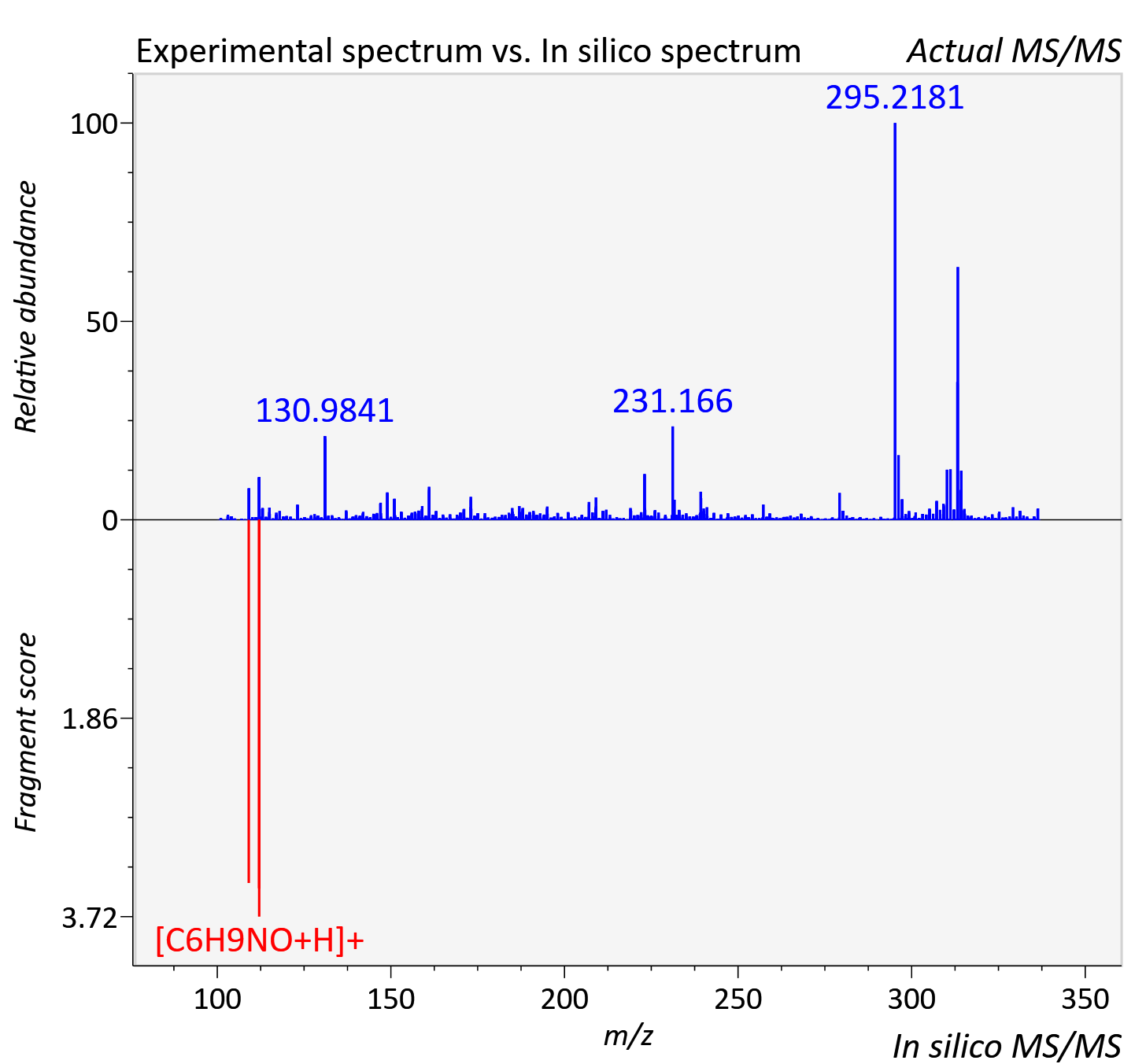
Experimental data
| Retention time | 12.8 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 336.327 | Theoretical mz | 336.326 |
| Error | 1.56 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.497 |
Identifiers and class information
| Inchi key | QQCGKIZHTJLRNN-JUJVUSMPSA-N |
|---|---|
| Smiles | O=C(C=CC=CCCCCCCCCCCCCC)NCC(C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| P21554 | CNR1 | Cannabinoid receptor 1 (by homology) | T76685 | SEA |
| P34913 | EPHX2 | Epoxide hydratase | T35734 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| Q8NER1 | TRPV1 | Vanilloid receptor | T83193 | SEA |
| Q99685 | MGLL | Monoglyceride lipase | T18664 | SEA |
| P34972 | CNR2 | Cannabinoid receptor 2 | T37693 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| Q9BXC1 | GPR174 | Probable G-protein coupled receptor 174 | T87661 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| Q9UP65 | PLA2G4C | Cytosolic phospholipase A2 gamma | T91113 | SEA |
| P40394 | ADH7 | Alcohol dehydrogenase class IV | T09770 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T76685 | DI0031 | Anorexia nervosa | [ICD-11: 6B80] | P21554 | CNR1 |
| T76685 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P21554 | CNR1 |
| T76685 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P21554 | CNR1 |
| T35734 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P34913 | EPHX2 |
| T35734 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P34913 | EPHX2 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T83193 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q8NER1 | TRPV1 |
| T18664 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q99685 | MGLL |
| T18664 | DI0409 | Tic disorder | [ICD-11: 8A05] | Q99685 | MGLL |
| T37693 | DI0040 | Attention deficit hyperactivity disorder | [ICD-11: 6A05] | P34972 | CNR2 |
| T37693 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P34972 | CNR2 |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
