Compound details
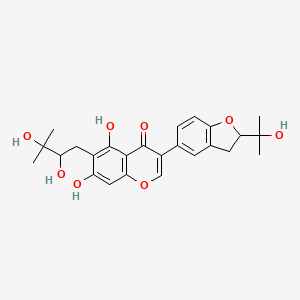
Lupinisoflavone M
| Compound ID | CDAMM02420 |
|---|---|
| Common name | Lupinisoflavone M | IUPAC name | 6-(2,3-dihydroxy-3-methylbutyl)-5,7-dihydroxy-3-[2-(2-hydroxypropan-2-yl)-2,3-dihydro-1-benzofuran-5-yl]chromen-4-one |
| Molecular formula | C25H28O8 |
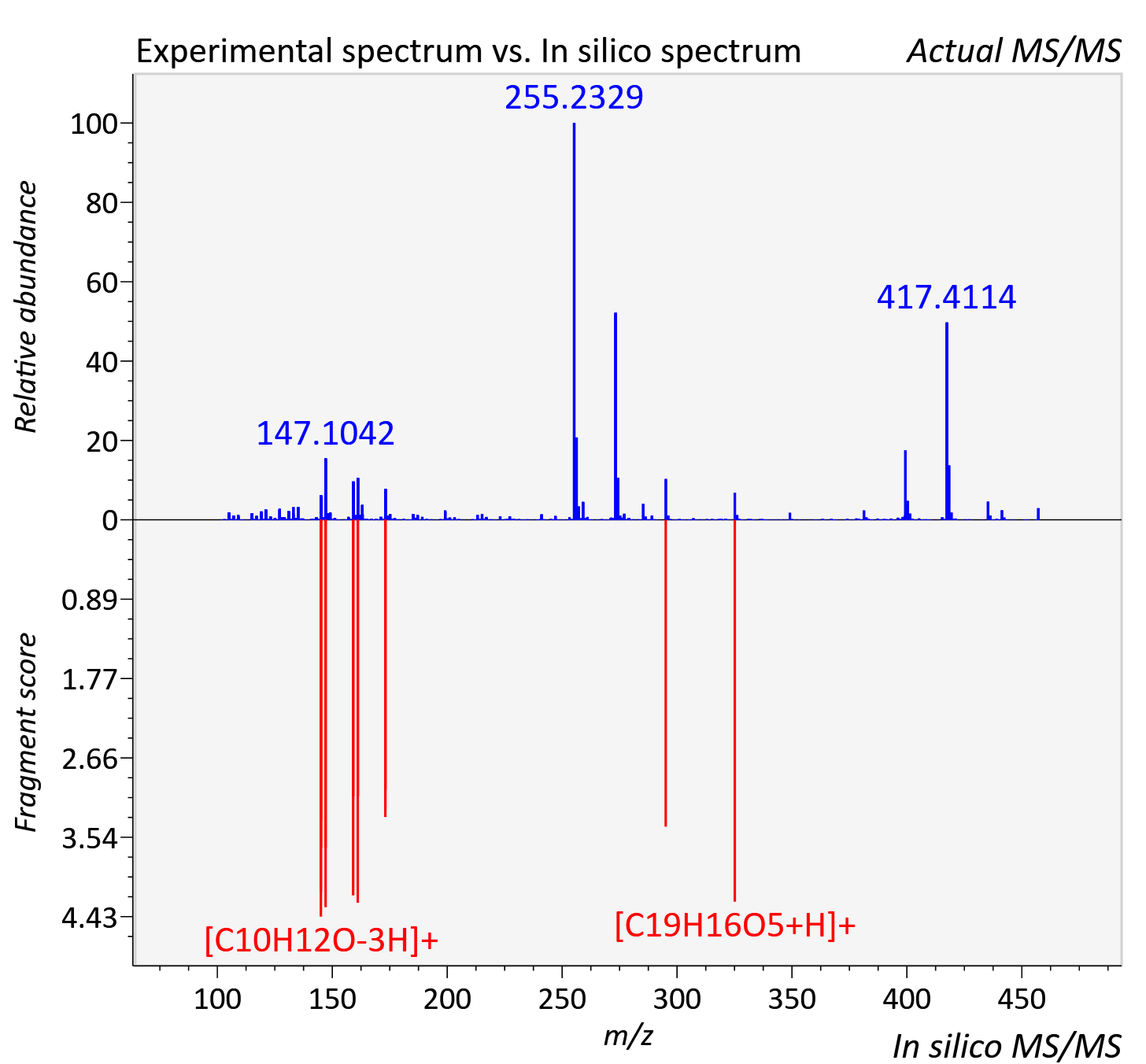
Experimental data
| Retention time | 7.77 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 457.185 | Theoretical mz | 457.185 |
| Error | 1.26 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.0393 |
Identifiers and class information
| Inchi key | IXEGNYHOJYAEHW-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1C(=COC2=CC(O)=C(C(O)=C12)CC(O)C(O)(C)C)C=3C=CC=4OC(CC4C3)C(O)(C)C |
| Superclass | Phenylpropanoids and polyketides |
| Class | Isoflavonoids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P13866 | SLC5A1 | Sodium/glucose cotransporter 1 | T54771 | SwissTargetPrediction |
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SwissTargetPrediction |
| P05121 | SERPINE1 | Plasminogen activator inhibitor-1 | T15556 | SwissTargetPrediction |
| P24723 | PRKCH | Protein kinase C eta | T40149 | SwissTargetPrediction |
| P05771 | PRKCB | Protein kinase C beta | T40276 | SwissTargetPrediction |
| Q02156 | PRKCE | Protein kinase C epsilon | T00895 | SwissTargetPrediction |
| Q05655 | PRKCD | Protein kinase C delta | T44861 | SwissTargetPrediction |
| P30837 | ALDH1B1 | Acetaldehyde dehydrogenase | T99641 | SwissTargetPrediction |
| P05129 | PRKCG | Protein kinase C gamma | T47107 | SwissTargetPrediction |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SwissTargetPrediction and SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| Q96HK3 | CALM | Calmodulin | T39610 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T54771 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | P13866 | SLC5A1 |
| T54771 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P13866 | SLC5A1 |
| T54771 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P13866 | SLC5A1 |
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T15556 | DI0037 | Asthma | [ICD-11: CA23] | P05121 | SERPINE1 |
| T15556 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | P05121 | SERPINE1 |
| T40149 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | P24723 | PRKCH |
| T40276 | DI0123 | Diffuse large B-cell lymphoma | [ICD-11: 2A81] | P05771 | PRKCB |
| T40276 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P05771 | PRKCB |
| T40276 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P05771 | PRKCB |
| T00895 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | Q02156 | PRKCE |
| T00895 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q02156 | PRKCE |
| T44861 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | Q05655 | PRKCD |
| T99641 | DI0396 | Substance abuse | [ICD-11: 6C40] | P30837 | ALDH1B1 |
| T47107 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P05129 | PRKCG |
| T47107 | DI0248 | Mastocytosis | [ICD-11: 2A21] | P05129 | PRKCG |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T39610 | DI0068 | Cardiac arrhythmia | [ICD-11: BC9Z] | Q96HK3 | CALM |
| T39610 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q96HK3 | CALM |
| T39610 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q96HK3 | CALM |
