Compound details
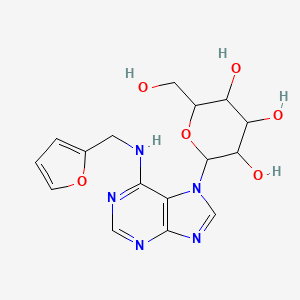
Kinetin-7-N-glucoside
| Compound ID | CDAMM02363 |
|---|---|
| Common name | Kinetin-7-N-glucoside | IUPAC name | 2-[6-(furan-2-ylmethylamino)purin-7-yl]-6-(hydroxymethyl)oxane-3,4,5-triol |
| Molecular formula | C16H19N5O6 |
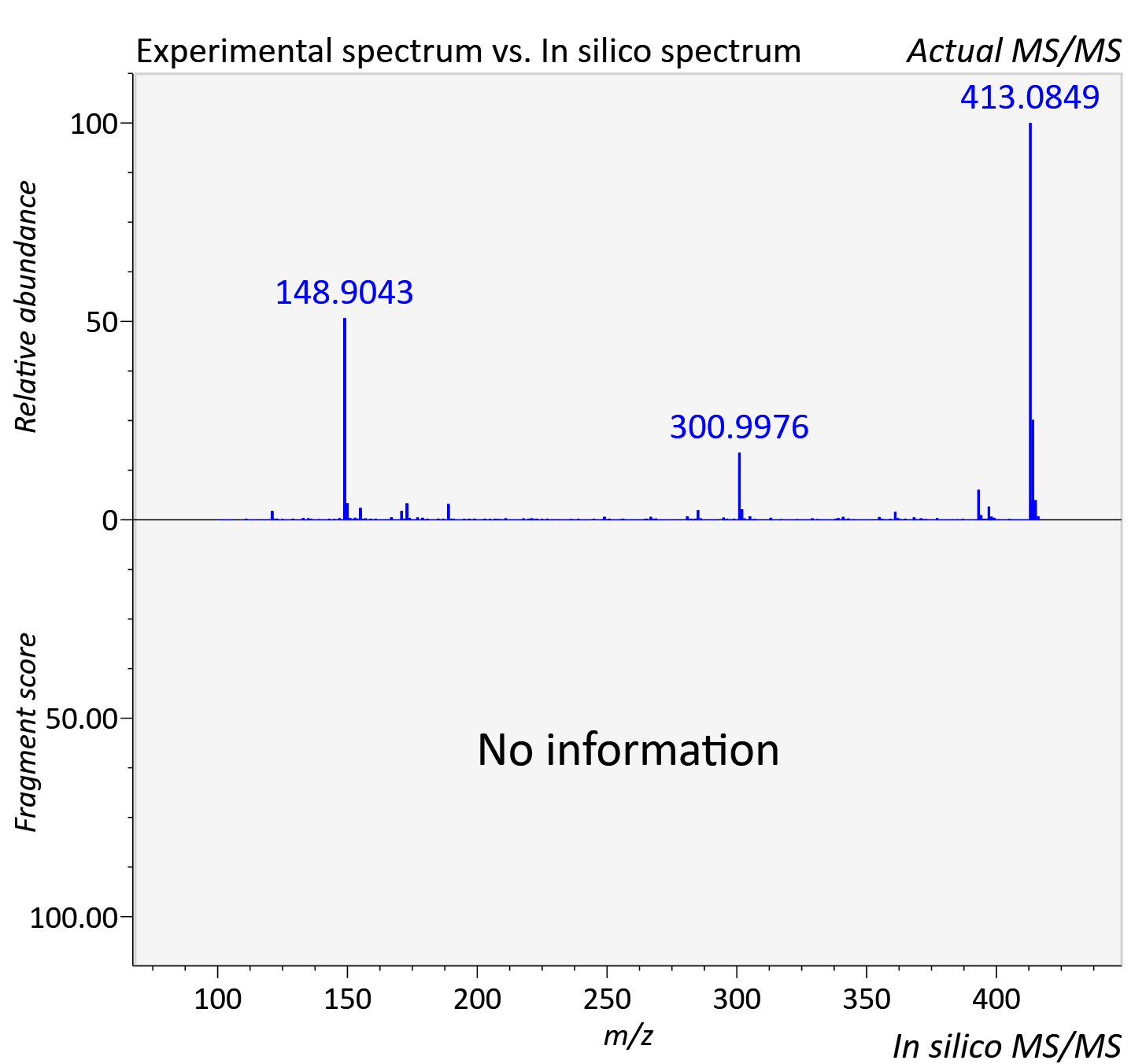
Experimental data
| Retention time | 14.79 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 416.097 | Theoretical mz | 416.097 |
| Error | 0.1 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.0862 |
Identifiers and class information
| Inchi key | AXJIOWGMASZFOU-HMXKMONRSA-N |
|---|---|
| Smiles | OCC1OC(N2C=NC=3N=CN=C(NCC=4OC=CC4)C32)C(O)C(O)C1O |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P29274 | ADORA2A | Adenosine A2a receptor | T77365 | SEA |
| P30542 | ADORA1 | Adenosine A1 receptor | T92072 | SEA |
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| P0DMS8 | ADORA3 | Adenosine A3 receptor | T36059 | SEA |
| P29275 | ADORA2B | Adenosine A2b receptor | T86679 | SEA |
| P26358 | DNMT1 | DNA (cytosine-5)-methyltransferase 1 (by homology) | T88304 | SEA |
| P32320 | CDA | Cytidine deaminase (by homology) | T79027 | SEA |
| P00813 | ADA | Adenosine deaminase | T03661 | SEA |
| P55263 | ADK | Adenosine kinase | T91661 | SEA |
| P04406 | GAPDH | Glyceraldehyde-3-phosphate dehydrogenase liver | T39321 | SEA |
| Q99808 | SLC29A1 | Equilibrative nucleoside transporter 1 | T13491 | SEA |
| O43868 | SLC28A2 | Sodium/nucleoside cotransporter 2 | T94627 | SEA |
| P23526 | AHCY | Adenosylhomocysteinase | T68698 | SEA |
| Q8TEK3 | DOT1L | Histone-lysine N-methyltransferase, H3 lysine-79 specific | T87686 | SEA |
| P55786 | NPEPPS | Puromycin-sensitive aminopeptidase | T15610 | SEA |
| P41231 | P2RY2 | Purinergic receptor P2Y2 | T93515 | SEA |
| Q96KQ7 | EHMT2 | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 | T53251 | SEA |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| Q9H9B1 | EHMT1 | Histone-lysine N-methyltransferase, H3 lysine-9 specific 5 | T39996 | SEA |
| Q9UBC3 | DNMT3B | DNA (cytosine-5)-methyltransferase 3B | T65501 | SEA |
| Q15391 | P2RY14 | Purinergic receptor P2Y14 | T16555 | SEA |
| P51575 | P2RX1 | P2X purinoceptor 1 | T69091 | SEA |
| P17707 | AMD1 | S-adenosylmethionine decarboxylase 1 | T55922 | SEA |
| Q8WTS6 | SETD7 | Histone-lysine N-methyltransferase SETD7 | T73184 | SEA |
| Q9NVM4 | PRMT7 | Protein arginine N-methyltransferase 7 | T79501 | SEA |
| O43463 | SUV39H1 | Histone-lysine N-methyltransferase SUV39H1 | T39519 | SEA |
| Q92800 | EZH1 | Histone-lysine N-methyltransferase EZH1 | T17986 | SEA |
| Q99571 | P2RX4 | P2X purinoceptor 4 | T60330 | SEA |
| Q13304 | GPR17 | Uracil nucleotide/cysteinyl leukotriene receptor | T33902 | SEA |
| Q86Y97 | KMT5C | Histone-lysine N-methyltransferase KMT5C | T68490 | SEA |
| Q9NZ70 | TAK1 | TGF-beta-activated kinase 1 | T04361 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T77365 | DI0319 | Orthostatic hypotension | [ICD-11: BA21] | P29274 | ADORA2A |
| T77365 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P29274 | ADORA2A |
| T77365 | DI0359 | Radionuclide imaging | [ICD-11: N.A.] | P29274 | ADORA2A |
| T92072 | DI0319 | Orthostatic hypotension | [ICD-11: BA21] | P30542 | ADORA1 |
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T36059 | DI0351 | Psoriasis | [ICD-11: EA90] | P0DMS8 | ADORA3 |
| T36059 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P0DMS8 | ADORA3 |
| T86679 | DI0180 | Herpes simplex infection | [ICD-11: 1F00] | P29275 | ADORA2B |
| T86679 | DI0397 | Supraventricular tachyarrhythmia | [ICD-11: BC81] | P29275 | ADORA2B |
| T88304 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P26358 | DNMT1 |
| T88304 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | P26358 | DNMT1 |
| T88304 | DI0286 | Myeloproliferative neoplasm | [ICD-11: 2A20] | P26358 | DNMT1 |
| T79027 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P32320 | CDA |
| T03661 | DI0016 | Adaptive immunity immunodeficiency | [ICD-11: 4A01] | P00813 | ADA |
| T03661 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P00813 | ADA |
| T03661 | DI0249 | Mature B-cell leukaemia | [ICD-11: 2A82] | P00813 | ADA |
| T91661 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P55263 | ADK |
| T39321 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P04406 | GAPDH |
| T39321 | DI0280 | Muscular dystrophy | [ICD-11: 8C70] | P04406 | GAPDH |
| T13491 | DI0030 | Angina pectoris | [ICD-11: BA40] | Q99808 | SLC29A1 |
| T87686 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | Q8TEK3 | DOT1L |
| T87686 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | Q8TEK3 | DOT1L |
| T87686 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | Q8TEK3 | DOT1L |
| T15610 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P55786 | NPEPPS |
| T15610 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | P55786 | NPEPPS |
| T93515 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P41231 | P2RY2 |
| T53251 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q96KQ7 | EHMT2 |
| T53251 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | Q96KQ7 | EHMT2 |
| T53251 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q96KQ7 | EHMT2 |
| T39996 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q9H9B1 | EHMT1 |
| T39996 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9H9B1 | EHMT1 |
| T65501 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9UBC3 | DNMT3B |
| T55922 | DI0004 | Acidosis | [ICD-11: 5C73] | P17707 | AMD1 |
| T73184 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q8WTS6 | SETD7 |
| T17986 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | Q92800 | EZH1 |
| T17986 | DI0238 | Lung cancer | [ICD-11: 2C25] | Q92800 | EZH1 |
| T17986 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | Q92800 | EZH1 |
| T17986 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | Q92800 | EZH1 |
| T17986 | DI0251 | Mature T-cell lymphoma | [ICD-11: 2A90] | Q92800 | EZH1 |
| T17986 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | Q92800 | EZH1 |
| T17986 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q92800 | EZH1 |
| T17986 | DI0423 | Ureteral cancer | [ICD-11: 2C92] | Q92800 | EZH1 |
| T04361 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | Q9NZ70 | TAK1 |
