Compound details
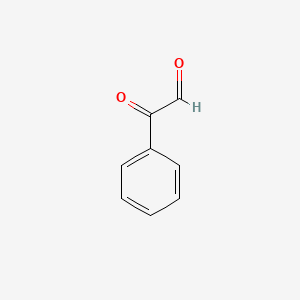
Phenylglyoxal
| Compound ID | CDAMM02335 |
|---|---|
| Common name | Phenylglyoxal | IUPAC name | 2-oxo-2-phenylacetaldehyde |
| Molecular formula | C8H6O2 |
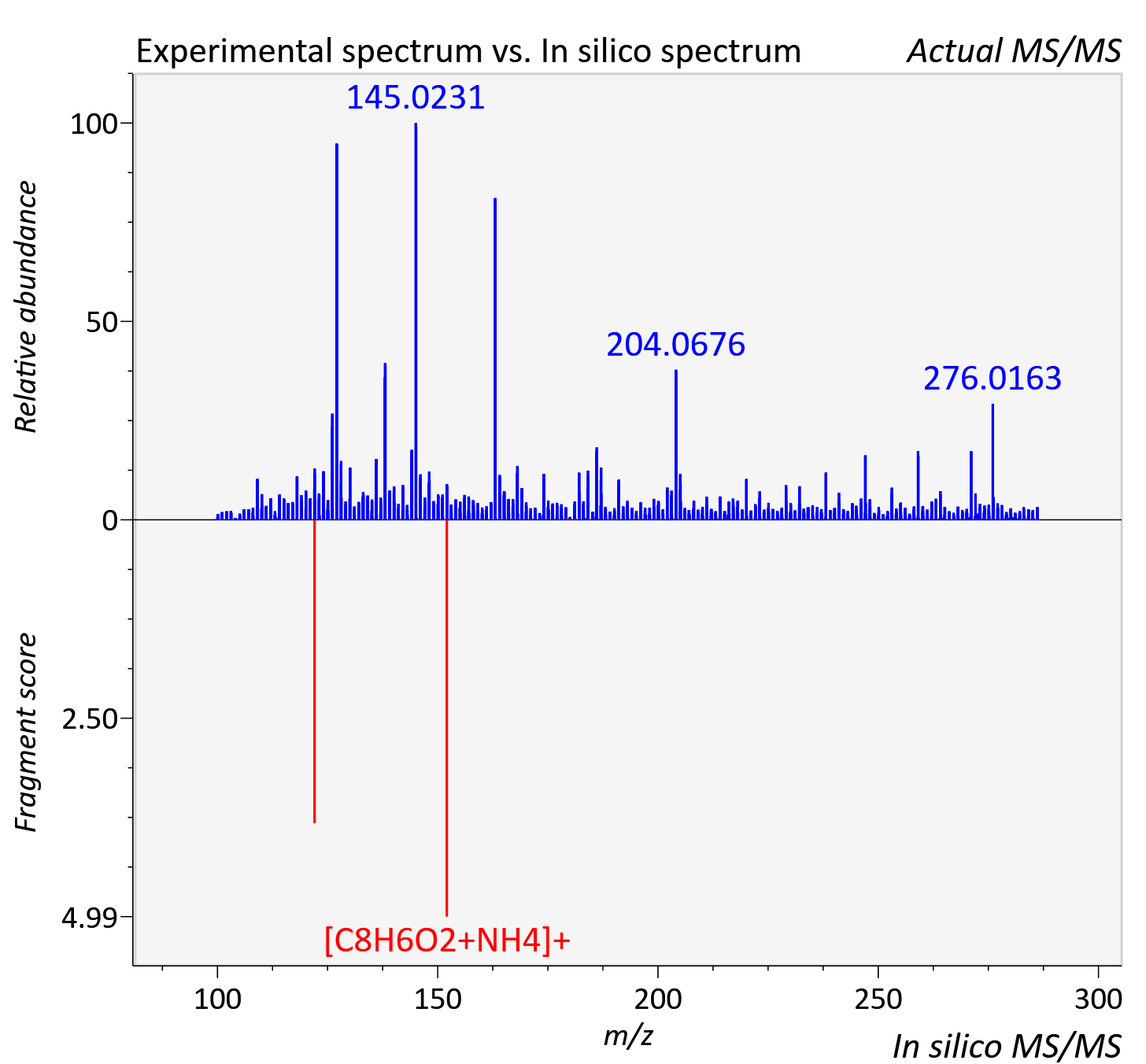
Experimental data
| Retention time | 0.43 |
|---|---|
| Adduct | [2M+NH4]+ |
| Actual mz | 286.106 | Theoretical mz | 286.108 |
| Error | 4.97 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.2813 |
Identifiers and class information
| Inchi key | OJUGVDODNPJEEC-UHFFFAOYSA-N |
|---|---|
| Smiles | O=CC(=O)C=1C=CC=CC1 |
| Superclass | Benzenoids |
| Class | Benzene and substituted derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P31213 | SRD5A2 | Steroid 5-alpha-reductase 2 | T71390 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| Q05940 | SLC18A2 | Synaptic vesicular amine transporter (by homology) | T48873 | SEA |
| Q9UQL6 | HDAC5 | Histone deacetylase 5 | T07930 | SEA |
| Q8WUI4 | HDAC7 | Histone deacetylase 7 | T98698 | SEA |
| Q9UKV0 | HDAC9 | Histone deacetylase 9 | T03687 | SEA |
| Q92769 | HDAC2 | Histone deacetylase 2 | T51191 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SwissTargetPrediction and SEA |
| P42330 | AKR1C3 | Aldo-keto-reductase family 1 member C3 | T60857 | SEA |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| P80404 | ABAT | Gamma-amino-N-butyrate transaminase | T59045 | SEA |
| Q9Y4C1 | KDM3A | Lysine-specific demethylase 3A | T25362 | SEA |
| P19838 | NFKB1 | Nuclear factor NF-kappa-B p105 subunit | T83145 | SEA |
| P08263 | GSTA1 | Glutathione S-transferase A1 | T17221 | SEA |
| P15085 | CPA1 | Carboxypeptidase A1 | T33966 | SEA |
| O14649 | KCNK3 | Potassium channel subfamily K member 3 | T15795 | SEA |
| Q9NPC2 | KCNK9 | Potassium channel subfamily K member 9 | T57613 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P19438 | TNFRSF1A | Tumor necrosis factor receptor R1 | T86552 | SEA |
| Q03405 | PLAUR | Urokinase plasminogen activator surface receptor | T74363 | SEA |
| P08311 | CTSG | Cathepsin G | T86385 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| O95551 | TDP2 | Tyrosyl-DNA phosphodiesterase 2 | T04696 | SEA |
| P51649 | ALDH5A1 | Succinate semialdehyde dehydrogenase | T13259 | SEA |
| O43175 | PHGDH | Phosphoglycerate dehydrogenase | T15139 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| Q9BY41 | HDAC8 | Histone deacetylase 8 | T28887 | SEA |
| P09172 | DBH | Dopamine beta hydroxylase | T74937 | SEA |
| P51843 | NR0B1 | Orphan nuclear receptor DAX-1 | T23191 | SEA |
| Q9C000 | NLRP1 | NLR pyrin domain containing 1 | T86910 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T71390 | DI0348 | Prostate hyperplasia | [ICD-11: GA90] | P31213 | SRD5A2 |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T48873 | DI0079 | Choreiform disorder | [ICD-11: 8A01] | Q05940 | SLC18A2 |
| T48873 | DI0125 | Dissociative neurological symptom disorder | [ICD-11: 6B60] | Q05940 | SLC18A2 |
| T48873 | DI0129 | Dystonic disorder | [ICD-11: 8A02] | Q05940 | SLC18A2 |
| T48873 | DI0137 | Essential hypertension | [ICD-11: BA00] | Q05940 | SLC18A2 |
| T48873 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q05940 | SLC18A2 |
| T51191 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q92769 | HDAC2 |
| T51191 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q92769 | HDAC2 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T60857 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P42330 | AKR1C3 |
| T59045 | DI0060 | Brain cancer | [ICD-11: 2A00] | P80404 | ABAT |
| T59045 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P80404 | ABAT |
| T59045 | DI0135 | Epileptic encephalopathy | [ICD-11: 8A62] | P80404 | ABAT |
| T59045 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P80404 | ABAT |
| T59045 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P80404 | ABAT |
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T15795 | DI0309 | Obstructive sleep apnoea | [ICD-11: 7A41] | O14649 | KCNK3 |
| T86552 | DI0060 | Brain cancer | [ICD-11: 2A00] | P19438 | TNFRSF1A |
| T86552 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P19438 | TNFRSF1A |
| T74363 | DI0286 | Myeloproliferative neoplasm | [ICD-11: 2A20] | Q03405 | PLAUR |
| T74363 | DI0404 | Thrombocytosis | [ICD-11: 3B63] | Q03405 | PLAUR |
| T86385 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P08311 | CTSG |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T13259 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P51649 | ALDH5A1 |
| T13259 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P51649 | ALDH5A1 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T28887 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9BY41 | HDAC8 |
| T74937 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P09172 | DBH |
| T74937 | DI0341 | Post-traumatic stress disorder | [ICD-11: 6B40] | P09172 | DBH |
