Compound details
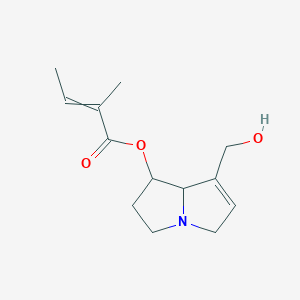
7-Angeloylheliotridine
| Compound ID | CDAMM02333 |
|---|---|
| Common name | 7-Angeloylheliotridine | IUPAC name | [7-(hydroxymethyl)-2,3,5,8-tetrahydro-1H-pyrrolizin-1-yl] 2-methylbut-2-enoate |
| Molecular formula | C13H19NO3 |
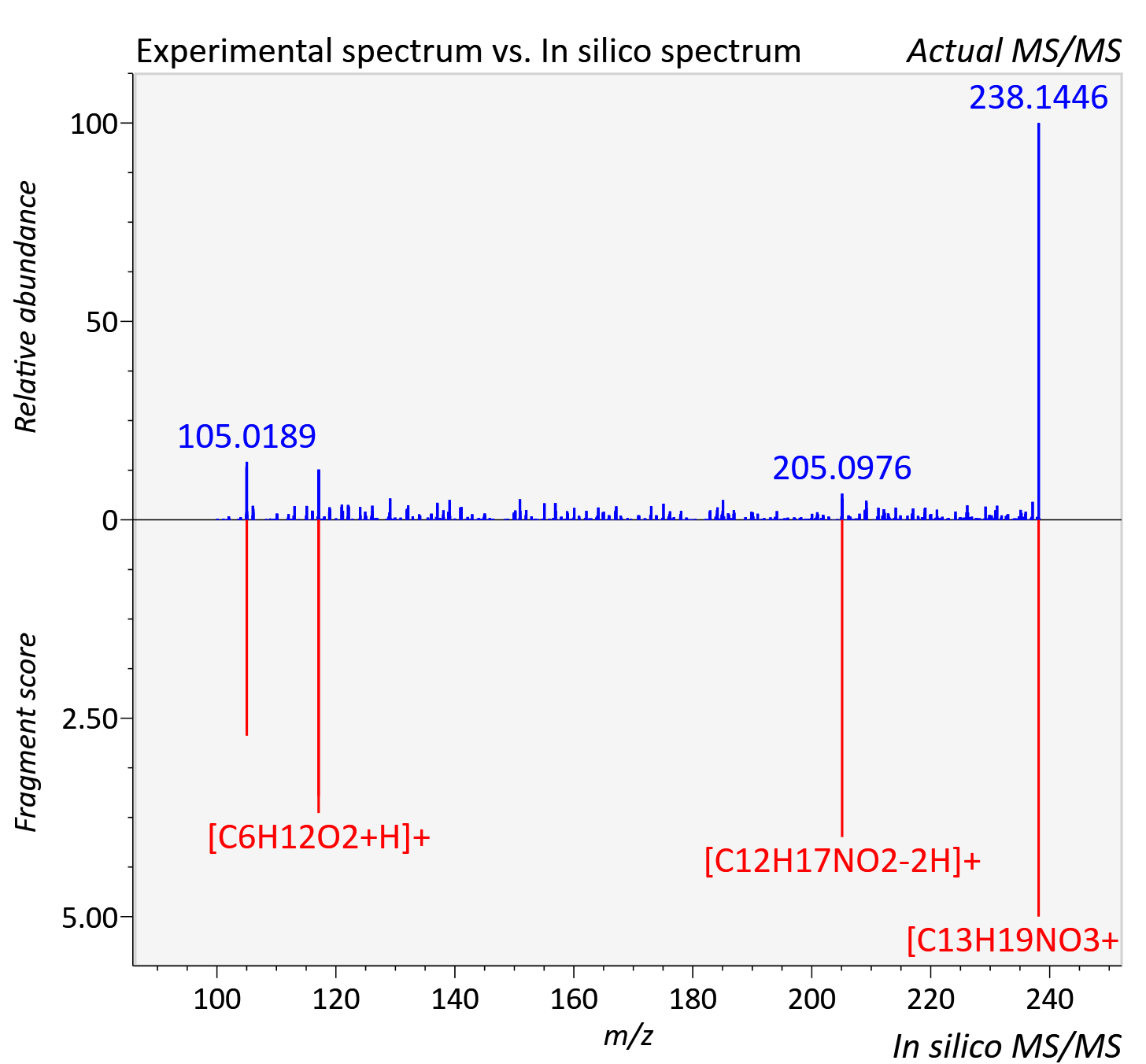
Experimental data
| Retention time | 9.43 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 238.144 | Theoretical mz | 238.143 |
| Error | 2.2 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.8922 |
Identifiers and class information
| Inchi key | TYGYPIIOOQNWBU-MCZAIVHYNA-N |
|---|---|
| Smiles | O=C(OC1CCN2CC=C(CO)C21)C(=CC)C |
| Superclass | Alkaloids and derivatives |
| Class |
Plant source
External sources
| HMDB ID | |
|---|---|
| KNApSAcK ID | C00002079 |
| FooDB ID | |
| DrugBank ID | |
| ChEBI ID | CHEBI:7666 |
| Pubchem Compound ID | 13457 |
| PlantCyc ID | |
| UNPD ID | UNPD112021 |
| Coconut ID | CNP0244343 |
| LipidMAPS ID | |
| NANPDB ID | NANPDB_873 |
