Compound details
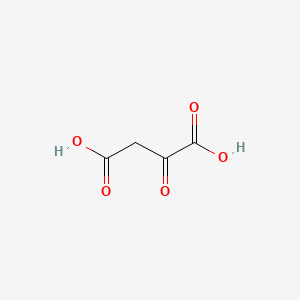
Oxalacetic acid
| Compound ID | CDAMM02320 |
|---|---|
| Common name | Oxalacetic acid | IUPAC name | 2-oxobutanedioic acid |
| Molecular formula | C4H4O5 |
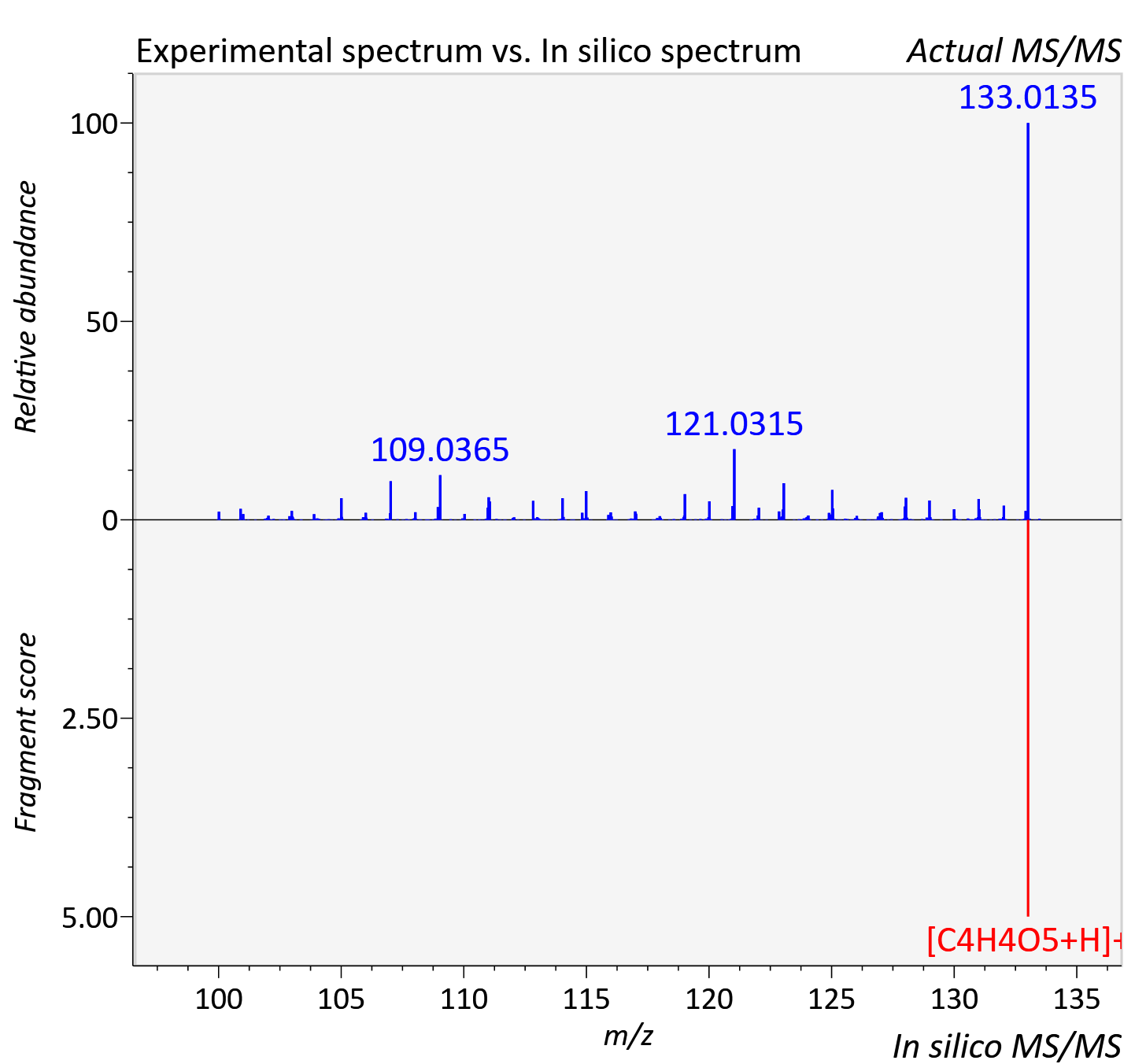
Experimental data
| Retention time | 15.12 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 133.014 | Theoretical mz | 133.013 |
| Error | 3.94 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.7874 |
Identifiers and class information
| Inchi key | KHPXUQMNIQBQEV-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C(O)C(=O)CC(=O)O |
| Superclass | Organic acids and derivatives |
| Class | Keto acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| Q9Y4C1 | KDM3A | Lysine-specific demethylase 3A | T25362 | SEA |
| O15054 | KDM6B | Lysine-specific demethylase 6B | T56057 | SEA |
| P41229 | KDM5C | Lysine-specific demethylase 5C | T85540 | SEA |
| Q9Y2K7 | KDM2A | Lysine-specific demethylase 2A | T71949 | SEA |
| Q9UPP1 | PHF8 | Histone lysine demethylase PHF8 | T00933 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
