Compound details
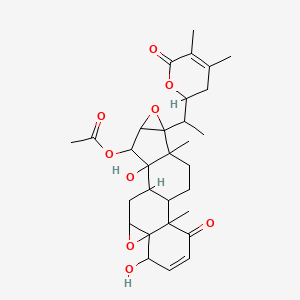
Physagulin C
| Compound ID | CDAMM02266 |
|---|---|
| Common name | Physagulin C | IUPAC name | [16-[1-(4,5-dimethyl-6-oxo-2,3-dihydropyran-2-yl)ethyl]-6,12-dihydroxy-2,17-dimethyl-3-oxo-8,15-dioxahexacyclo[9.8.0.02,7.07,9.012,17.014,16]nonadec-4-en-13-yl] acetate |
| Molecular formula | C30H38O9 |
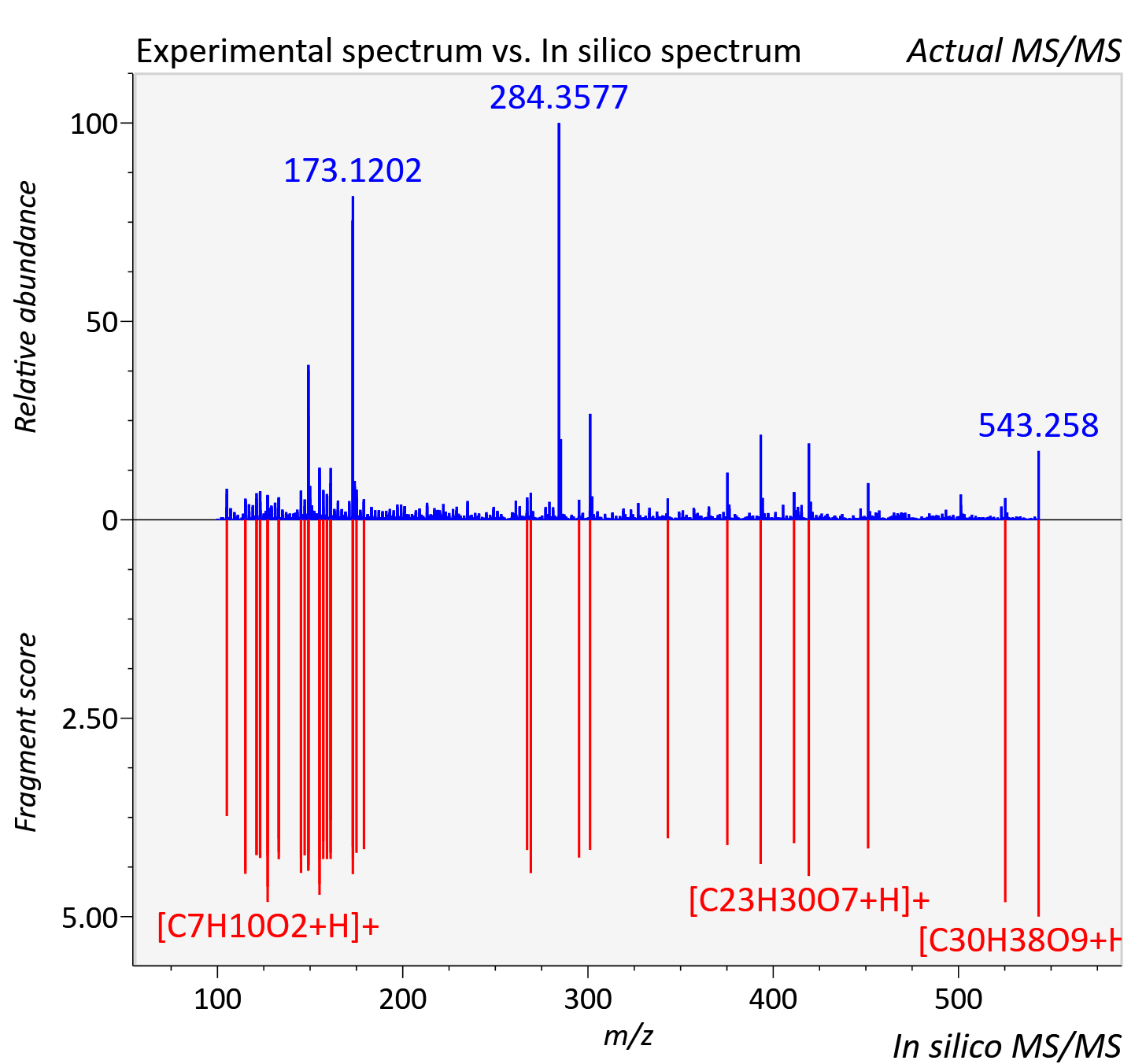
Experimental data
| Retention time | 10.42 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 543.26 | Theoretical mz | 543.259 |
| Error | 0.41 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.8926 |
Identifiers and class information
| Inchi key | VVQRJFUYBNCTQX-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC(CC(=C1C)C)C(C)C23OC3C(OC(=O)C)C4(O)C5CC6OC76C(O)C=CC(=O)C7(C)C5CCC42C |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SwissTargetPrediction |
| P35354 | PTGS2 | Cyclooxygenase-2 | T66665 | SwissTargetPrediction |
| P10275 | AR | Androgen Receptor | T11211 | SwissTargetPrediction |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction |
| O14746 | TERT | Telomerase reverse transcriptase | T86052 | SwissTargetPrediction |
| Q08499 | PDE4D | Phosphodiesterase 4D | T02001 | SwissTargetPrediction |
| Q02156 | PRKCE | Protein kinase C epsilon | T00895 | SwissTargetPrediction |
| Q05655 | PRKCD | Protein kinase C delta | T44861 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T66665 | DI0087 | Chronic pain | [ICD-11: MG30] | P35354 | PTGS2 |
| T66665 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P35354 | PTGS2 |
| T66665 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P35354 | PTGS2 |
| T66665 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P35354 | PTGS2 |
| T66665 | DI0227 | Knee osteoarthritis | [ICD-11: FA01] | P35354 | PTGS2 |
| T66665 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35354 | PTGS2 |
| T66665 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35354 | PTGS2 |
| T66665 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P35354 | PTGS2 |
| T66665 | DI0416 | Tuberculosis | [ICD-11: 1B10-1B12] | P35354 | PTGS2 |
| T11211 | DI0005 | Acne vulgaris | [ICD-11: ED80] | P10275 | AR |
| T11211 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P10275 | AR |
| T11211 | DI0021 | Alcoholic liver disease | [ICD-11: DB94] | P10275 | AR |
| T11211 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P10275 | AR |
| T11211 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P10275 | AR |
| T11211 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P10275 | AR |
| T11211 | DI0194 | Hypo-osmolality/hyponatraemia | [ICD-11: 5C72] | P10275 | AR |
| T11211 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | P10275 | AR |
| T11211 | DI0247 | Mammary tumour | [ICD-11: 2C60-2C6Z] | P10275 | AR |
| T11211 | DI0346 | Prostate cancer | [ICD-11: 2C82] | P10275 | AR |
| T11211 | DI0401 | Testicular dysfunction | [ICD-11: 5A81] | P10275 | AR |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T86052 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | O14746 | TERT |
| T86052 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | O14746 | TERT |
| T86052 | DI0326 | Pancreatic cancer | [ICD-11: 2C10] | O14746 | TERT |
| T02001 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | Q08499 | PDE4D |
| T00895 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | Q02156 | PRKCE |
| T00895 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q02156 | PRKCE |
| T44861 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | Q05655 | PRKCD |
