Compound details
Jioglutolide
| Compound ID | CDAMM02185 |
|---|---|
| Common name | Jioglutolide | IUPAC name | 5,7-dihydroxy-7-methyl-1,4,4a,5,6,7a-hexahydrocyclopenta[c]pyran-3-one |
| Molecular formula | C9H14O4 |
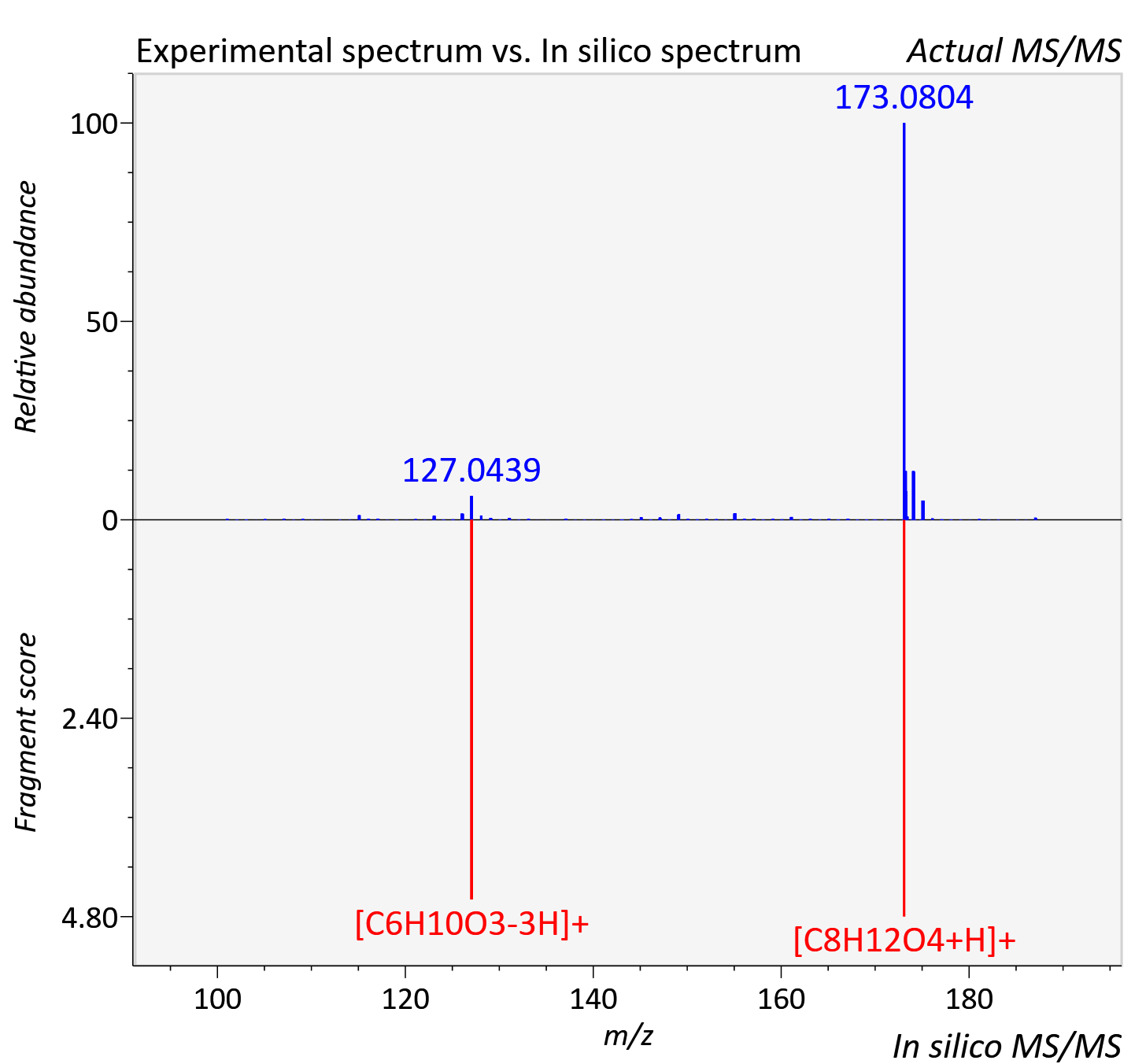
Experimental data
| Retention time | 10.31 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 187.097 | Theoretical mz | 187.096 |
| Error | 1.73 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.0721 |
Identifiers and class information
| Inchi key | LKZXAOPEOQRJLJ-IFQPIYTINA-N |
|---|---|
| Smiles | O=C1OCC2C(C1)C(O)CC2(O)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
External sources
| HMDB ID | |
|---|---|
| KNApSAcK ID | C00010585 |
| FooDB ID | |
| DrugBank ID | |
| ChEBI ID | |
| Pubchem Compound ID | 14413762 |
| PlantCyc ID | |
| UNPD ID | UNPD48464 |
| Coconut ID | CNP0180136 |
| LipidMAPS ID | |
| NANPDB ID | NANPDB_789 |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P17612 | PRKACA | cAMP-dependent protein kinase alpha-catalytic subunit | T12808 | SwissTargetPrediction |
| P04035 | HMGCR | HMG-CoA reductase | T53585 | SwissTargetPrediction |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T12808 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P17612 | PRKACA |
| T53585 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P04035 | HMGCR |
| T53585 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P04035 | HMGCR |
| T53585 | DI0128 | Dyslipidemia | [ICD-11: 5C80-5C81] | P04035 | HMGCR |
| T53585 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P04035 | HMGCR |
| T53585 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P04035 | HMGCR |
| T53585 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P04035 | HMGCR |
| T53585 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P04035 | HMGCR |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
