Compound details
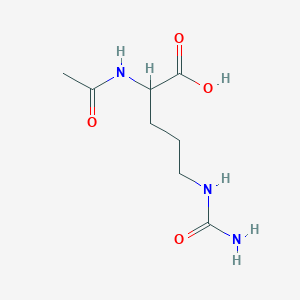
N-a-Acetylcitrulline
| Compound ID | CDAMM02174 |
|---|---|
| Common name | N-a-Acetylcitrulline | IUPAC name | 2-acetamido-5-(carbamoylamino)pentanoic acid |
| Molecular formula | C8H15N3O4 |
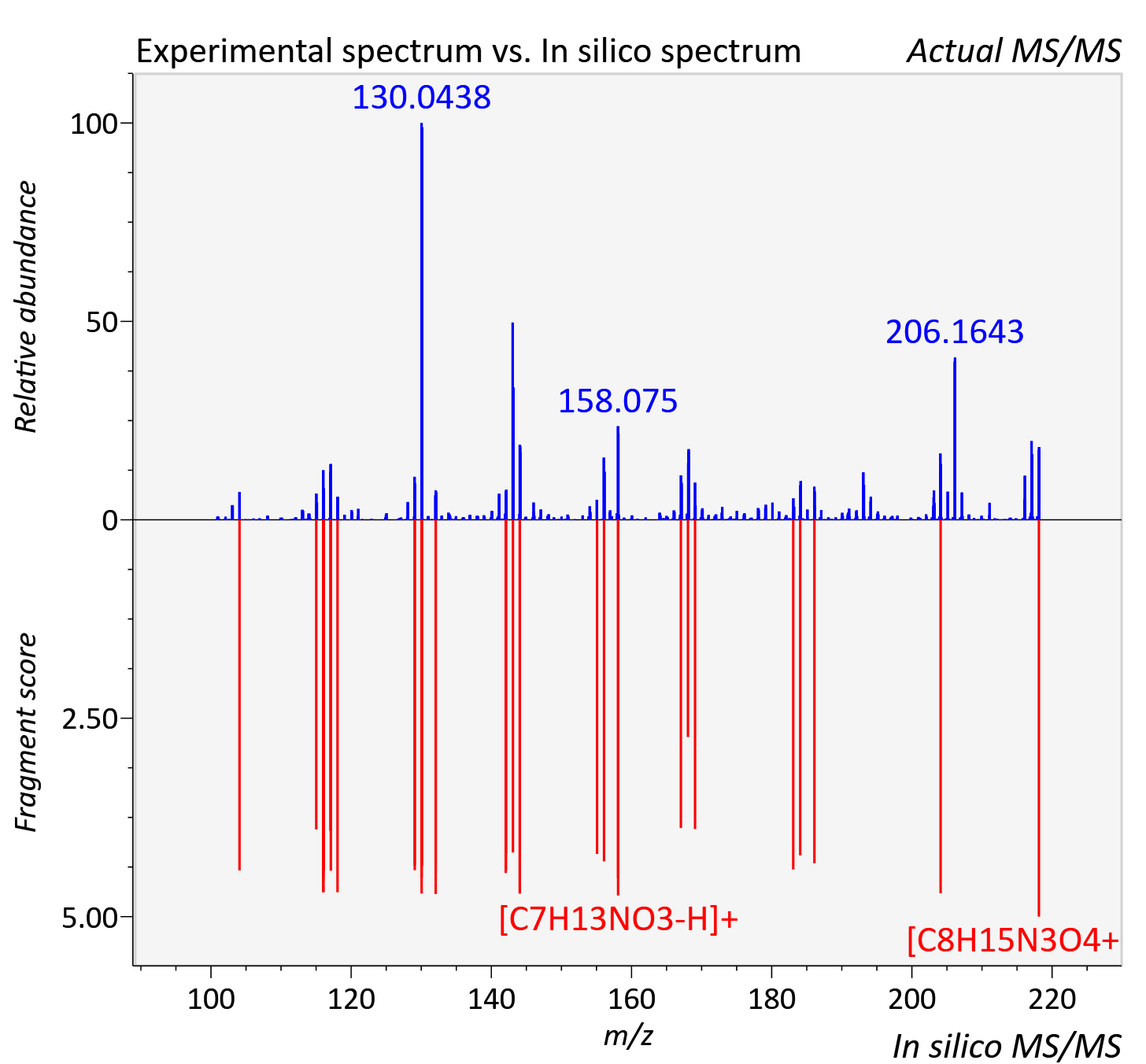
Experimental data
| Retention time | 8.6 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 218.114 | Theoretical mz | 218.113 |
| Error | 4.69 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.0663 |
Identifiers and class information
| Inchi key | WMQMIOYQXNRROC-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(N)NCCCC(NC(=O)C)C(=O)O |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P29475 | NOS1 | Nitric-oxide synthase, brain | T16117 | SEA |
| P35228 | NOS2 | Nitric oxide synthase, inducible | T02703 | SEA |
| P29474 | NOS3 | Nitric-oxide synthase, endothelial | T06046 | SEA |
| Q9NTG7 | SIRT3 | NAD-dependent deacetylase sirtuin 3 | T91959 | SEA |
| Q8IXJ6 | SIRT2 | NAD-dependent deacetylase sirtuin 2 | T83904 | SEA |
| P08514 | ITGA2B | Integrin alpha-IIb/beta-3 | T50688 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| Q96IY4 | CPB2 | Carboxypeptidase B2 isoform A | T98022 | SEA |
| P17706 | PTPN2 | T-cell protein-tyrosine phosphatase | T49156 | SEA |
| P15086 | CPB1 | Carboxypeptidase B | T36725 | SEA |
| P32245 | MC4R | Melanocortin receptor 4 | T72458 | SEA |
| Q9ULW8 | PADI3 | Protein-arginine deiminase type-3 | T36612 | SEA |
| Q9UM07 | PADI4 | Protein-arginine deiminase type-4 | T50429 | SEA |
| P25929 | NPY1R | Neuropeptide Y receptor type 1 | T89213 | SEA |
| Q16581 | C3AR1 | C3a anaphylatoxin chemotactic receptor | T30219 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SEA |
| Q16549 | PCSK7 | Subtilisin/kexin type 7 | T33211 | SEA |
| Q9UIQ6 | LNPEP | Cystinyl aminopeptidase | T97537 | SEA |
| P61964 | WDR5 | WD repeat-containing protein 5 | T77393 | SEA |
| Q9Y5Y6 | ST14 | Matriptase | T31543 | SEA |
| O95931 | CBX7 | Chromobox protein homolog 7 | T72319 | SEA |
| Q04756 | HGFAC | Hepatocyte growth factor activator | T38584 | SEA |
| P09668 | CTSH | Cathepsin (H and K) | T40103 | SEA |
| O60235 | TMPRSS11D | Transmembrane protease serine 11D | T00302 | SEA |
| O14786 | NRP1 | Neuropilin-1 (by homology) | T15023 | SEA |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SEA |
| P09958 | FUR | HUMAN dibasic-processing enzyme | T98945 | SEA |
| P29120 | PCSK1 | Neuroendocrine convertase 1 | T36445 | SEA |
| Q92824 | PCSK5 | Proprotein convertase subtilisin/kexin type 5 | T38581 | SEA |
| Q9GZQ6 | NPFFR1 | Neuropeptide FF receptor 1 | T79780 | SEA |
| Q9Y5X5 | NPFFR2 | Neuropeptide FF receptor 2 | T90818 | SEA |
| O60259 | KLK8 | Neuropsin | T43289 | SEA |
| Q9Y5K2 | KLK4 | Kallikrein-4 | T79400 | SEA |
| Q9Y4G9 | PCSK6 | Proprotein convertase subtilisin/kexin type 6 | T86503 | SEA |
| P17342 | NPR3 | Natriuretic peptide receptor | T92963 | SEA |
| P16066 | NPR1 | Atrial natriuretic peptide receptor A | T71230 | SEA |
| P05106 | ITGB3 | Integrin beta-3 | T26457 | SEA |
| P00480 | OTC | Ornithine transcarbamylase | T52098 | SEA |
| Q16348 | PEPT2 | Solute carrier family 15 member 2 | T52067 | SEA |
| Q9NXA8 | SIRT5 | NAD-dependent deacetylase sirtuin-5 | T91940 | SEA |
| Q9BX92 | CFB | Complement factor B | T66011 | SEA |
| P05981 | HPN | Serine protease hepsin | T73471 | SEA |
| P16519 | PCSK2 | Neuroendocrine convertase 2 | T97749 | SEA |
| P50391 | NPY4R | Neuropeptide Y receptor type 4 | T27944 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16117 | DI0173 | Headache | [ICD-11: 8A80-8A84] | P29475 | NOS1 |
| T16117 | DI0264 | Migraine | [ICD-11: 8A80] | P29475 | NOS1 |
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T06046 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P29474 | NOS3 |
| T83904 | DI0365 | Retinopathy | [ICD-11: 9B71] | Q8IXJ6 | SIRT2 |
| T50688 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P08514 | ITGA2B |
| T50688 | DI0030 | Angina pectoris | [ICD-11: BA40] | P08514 | ITGA2B |
| T50688 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P08514 | ITGA2B |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T98022 | DI0073 | Cerebral ischaemia | [ICD-11: 8B1Z] | Q96IY4 | CPB2 |
| T98022 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | Q96IY4 | CPB2 |
| T98022 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | Q96IY4 | CPB2 |
| T49156 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P17706 | PTPN2 |
| T72458 | DI0192 | Hypoactive sexual desire dysfunction | [ICD-11: HA00] | P32245 | MC4R |
| T72458 | DI0228 | Large intestine motility disorder | [ICD-11: DB32] | P32245 | MC4R |
| T72458 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P32245 | MC4R |
| T89213 | DI0050 | Binge eating disorder | [ICD-11: 6B82] | P25929 | NPY1R |
| T89213 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P25929 | NPY1R |
| T89213 | DI0174 | Heart disease | [ICD-11: BA41-BA42] | P25929 | NPY1R |
| T89213 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P25929 | NPY1R |
| T89213 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P25929 | NPY1R |
| T15023 | DI0365 | Retinopathy | [ICD-11: 9B71] | O14786 | NRP1 |
| T97071 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | Q04609 | FOLH1 |
| T97071 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q04609 | FOLH1 |
| T98945 | DI0263 | Middle East Respiratory Syndrome | [ICD-11: 1D64] | P09958 | FUR |
| T92963 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P17342 | NPR3 |
| T71230 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P16066 | NPR1 |
| T52098 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P00480 | OTC |
| T66011 | DI0170 | Haemolytic anemia | [ICD-11: 3A20-3A2Z] | Q9BX92 | CFB |
| T27944 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P50391 | NPY4R |
| T27944 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P50391 | NPY4R |
| T27944 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P50391 | NPY4R |
