Compound details
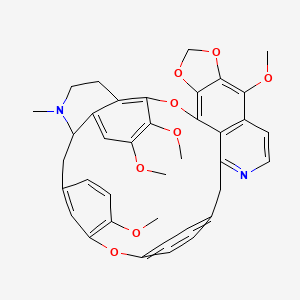
Thalfine
| Compound ID | CDAMM02166 |
|---|---|
| Common name | Thalfine | IUPAC name | 8,17,18,30-tetramethoxy-24-methyl-10,12,15,32-tetraoxa-4,24-diazaoctacyclo[31.2.2.13,7.127,31.09,13.016,21.020,25.014,39]nonatriaconta-1(35),3,5,7(39),8,13,16(21),17,19,27(38),28,30,33,36-tetradecaene |
| Molecular formula | C38H36N2O8 |
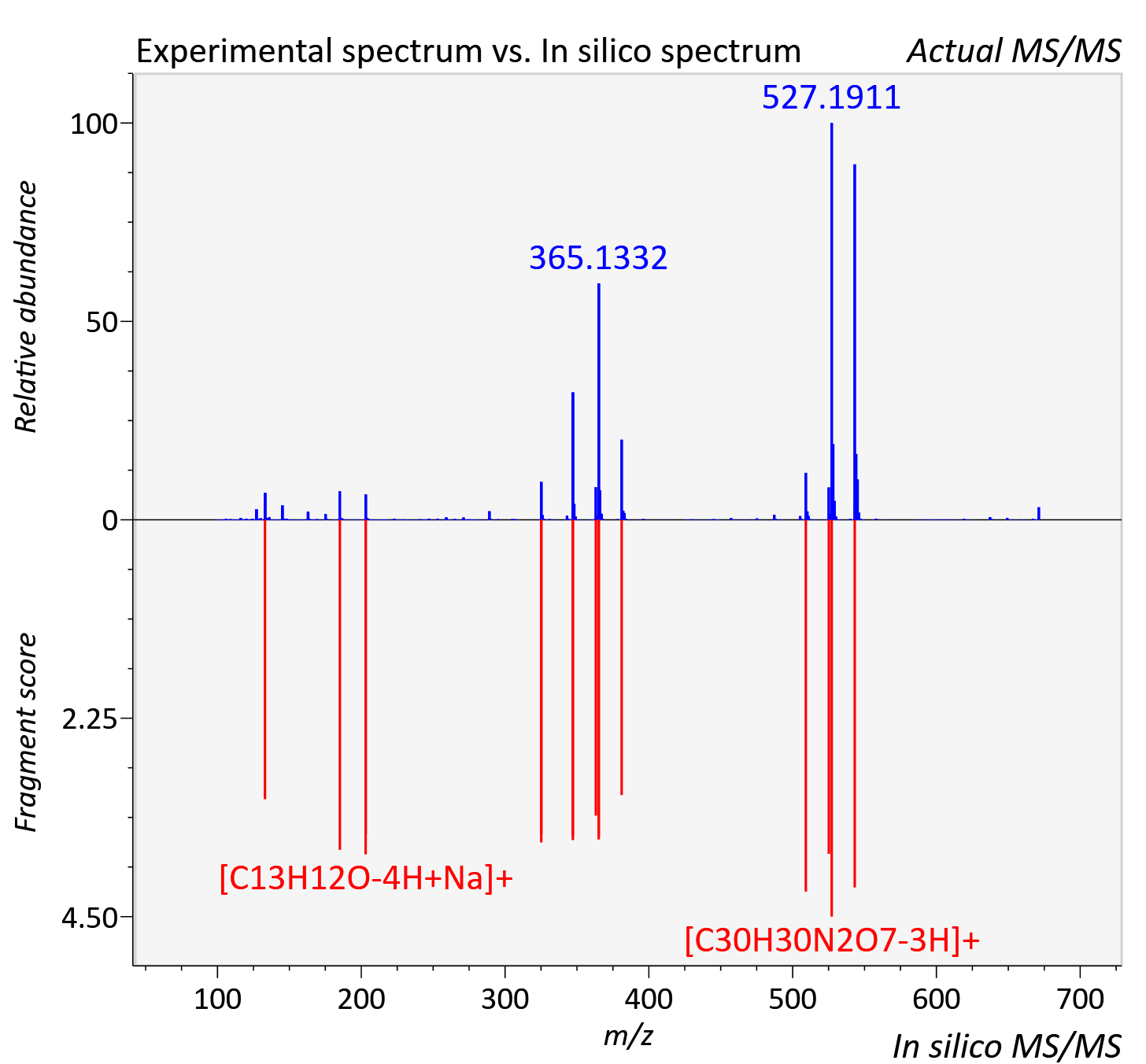
Experimental data
| Retention time | 0.43 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 671.237 | Theoretical mz | 671.236 |
| Error | 0.57 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.6903 |
Identifiers and class information
| Inchi key | OSOKLEWJPLGVBW-DKKPLQGMNA-N |
|---|---|
| Smiles | N=1C=CC=2C(OC)=C3OCOC3=C4OC=5C(OC)=C(OC)C=C6C5CCN(C)C6CC7=CC=C(OC)C(OC8=CC=C(C=C8)CC1C42)=C7 |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T55815 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | Q15822 | CHRNA2 |
| T55815 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | Q15822 | CHRNA2 |
| T55815 | DI0166 | Glaucoma | [ICD-11: 9C61] | Q15822 | CHRNA2 |
| T55815 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | Q15822 | CHRNA2 |
| T55815 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | Q15822 | CHRNA2 |
