Compound details
5,7alpha-Dihydro-1,4,4,7a-tetramethyl-4H-indene
| Compound ID | CDAMM02142 |
|---|---|
| Common name | 5,7alpha-Dihydro-1,4,4,7a-tetramethyl-4H-indene | IUPAC name | 1,4,4,7a-tetramethyl-5H-indene |
| Molecular formula | C13H18 |
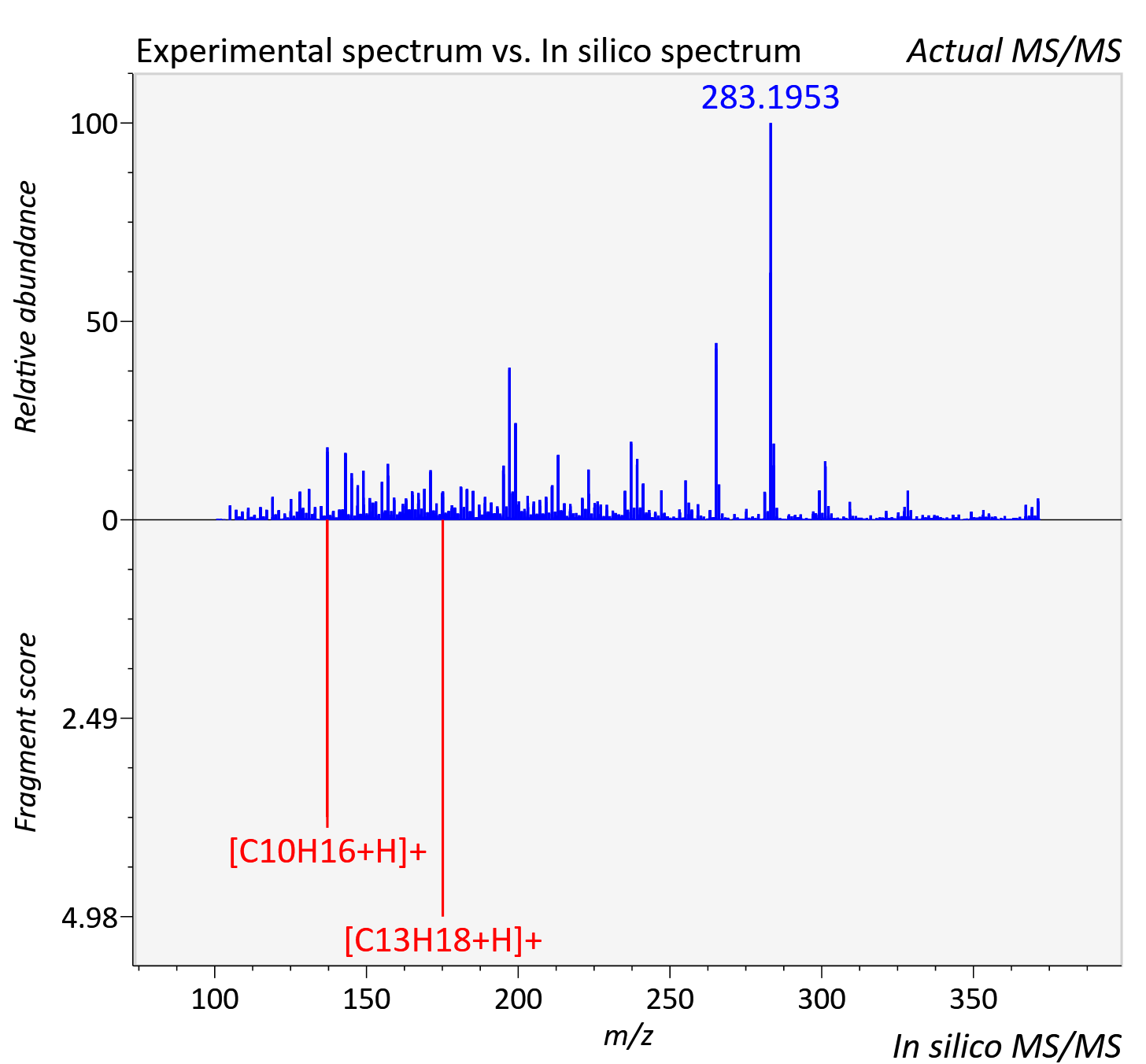
Experimental data
| Retention time | 8.9 |
|---|---|
| Adduct | [2M+Na]+ |
| Actual mz | 371.272 | Theoretical mz | 371.271 |
| Error | 1.57 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.1536 |
Identifiers and class information
| Inchi key | XOFDOXJFXCEFDE-UHFFFAOYNA-N |
|---|---|
| Smiles | C=1C=C2C(C=CCC2(C)C)(C1C)C |
| Superclass | Hydrocarbons |
| Class | Unsaturated hydrocarbons |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T77365 | DI0319 | Orthostatic hypotension | [ICD-11: BA21] | P29274 | ADORA2A |
| T77365 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P29274 | ADORA2A |
| T77365 | DI0359 | Radionuclide imaging | [ICD-11: N.A.] | P29274 | ADORA2A |
| T92072 | DI0319 | Orthostatic hypotension | [ICD-11: BA21] | P30542 | ADORA1 |
| T36059 | DI0351 | Psoriasis | [ICD-11: EA90] | P0DMS8 | ADORA3 |
| T36059 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P0DMS8 | ADORA3 |
