Compound details
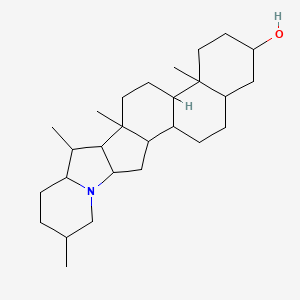
3-Epidemissidine
| Compound ID | CDAMM02131 |
|---|---|
| Common name | 3-Epidemissidine | IUPAC name | 10,14,16,20-tetramethyl-22-azahexacyclo[12.10.0.02,11.05,10.015,23.017,22]tetracosan-7-ol |
| Molecular formula | C27H45NO |
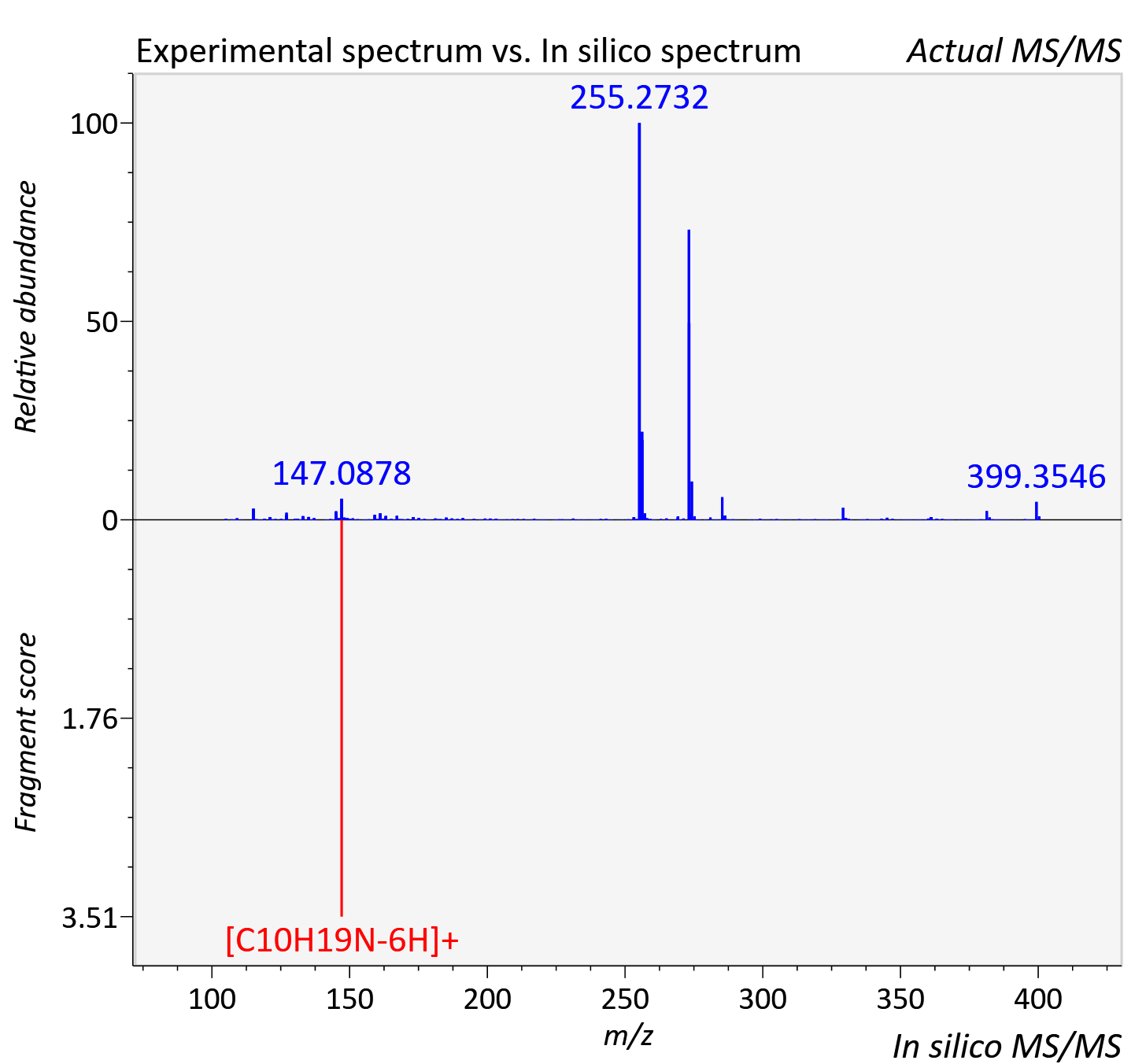
Experimental data
| Retention time | 4.97 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 400.357 | Theoretical mz | 400.357 |
| Error | 0.69 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.3665 |
Identifiers and class information
| Inchi key | JALVTHFTYRPDMB-UHFFFAOYNA-N |
|---|---|
| Smiles | OC1CCC2(C)C(CCC3C2CCC4(C)C3CC5N6CC(C)CCC6C(C)C54)C1 |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q99720 | SIGMAR1 | Sigma opioid receptor | T46360 | SwissTargetPrediction |
| P00915 | CA1 | Carbonic anhydrase I | T13201 | SEA |
| P00918 | CA2 | Carbonic anhydrase II | T20401 | SEA |
| Q15125 | EBP | Anti-estrogen binding site (AEBS) | T25317 | SwissTargetPrediction and SEA |
| Q05586 | GRIN1 | Glutamate NMDA receptor; GRIN1/GRIN2B | T62276 | SEA |
| P08185 | SERPINA6 | Corticosteroid binding globulin | T88452 | SEA |
| Q8TDU6 | GPBAR1 | G-protein coupled bile acid receptor 1 | T86273 | SEA |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
| P60842 | EIF4A1 | Eukaryotic initiation factor 4A-I | T86805 | SEA |
| Q13224 | GRIN2B | Glutamate [NMDA] receptor subunit epsilon 2 | T76414 | SEA |
| O15399 | GRIN2D | Glutamate [NMDA] receptor subunit epsilon 4 | T56588 | SEA |
| Q14973 | SLC10A1 | Bile acid transporter | T99189 | SEA |
| P38571 | LIPA | Lysosomal acid lipase/cholesteryl ester hydrolase | T24373 | SEA |
| P47870 | GABRB2 | GABA(A) receptor beta-2 | T80387 | SEA |
| O15439 | ABCC4 | Multidrug resistance protein 4 | T39919 | SEA |
| Q14957 | GRIN2C | Glutamate receptor ionotropic NMDA 2C | T72595 | SEA |
| O14764 | GABRD | GABA(A) receptor delta | T27376 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T46360 | DI0105 | Cough | [ICD-11: MD12] | Q99720 | SIGMAR1 |
| T13201 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00915 | CA1 |
| T13201 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00915 | CA1 |
| T20401 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P00918 | CA2 |
| T20401 | DI0137 | Essential hypertension | [ICD-11: BA00] | P00918 | CA2 |
| T20401 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00918 | CA2 |
| T20401 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P00918 | CA2 |
| T20401 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P00918 | CA2 |
| T20401 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00918 | CA2 |
| T25317 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q15125 | EBP |
| T62276 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q05586 | GRIN1 |
| T62276 | DI0182 | HIV-infected patients with tuberculosis | [ICD-11: 1B10-1B14] | Q05586 | GRIN1 |
| T88452 | DI0037 | Asthma | [ICD-11: CA23] | P08185 | SERPINA6 |
| T88452 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P08185 | SERPINA6 |
| T88452 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08185 | SERPINA6 |
| T86273 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | Q8TDU6 | GPBAR1 |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T86805 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P60842 | EIF4A1 |
| T76414 | DI0024 | Alpha-1-antitrypsin deficiency | [ICD-11: 5C5A] | Q13224 | GRIN2B |
| T56588 | DI0296 | Neurodegenerative disorder | [ICD-11: 8A20-8A23] | O15399 | GRIN2D |
| T99189 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | Q14973 | SLC10A1 |
| T24373 | DI0132 | Enzyme deficiency | [ICD-11: 5C51-5C57] | P38571 | LIPA |
| T80387 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P47870 | GABRB2 |
| T80387 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P47870 | GABRB2 |
| T80387 | DI0256 | Mental/behavioural/neurodevelopmental disorder | [ICD-11: 6E20-6E8Z] | P47870 | GABRB2 |
| T80387 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P47870 | GABRB2 |
| T72595 | DI0296 | Neurodegenerative disorder | [ICD-11: 8A20-8A23] | Q14957 | GRIN2C |
| T27376 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O14764 | GABRD |
| T27376 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | O14764 | GABRD |
| T27376 | DI0256 | Mental/behavioural/neurodevelopmental disorder | [ICD-11: 6E20-6E8Z] | O14764 | GABRD |
