Compound details
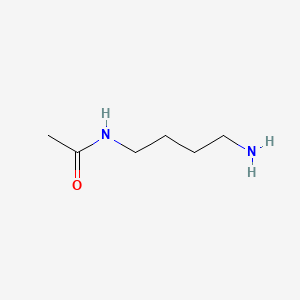
N-Acetylputrescine
| Compound ID | CDAMM02120 |
|---|---|
| Common name | N-Acetylputrescine | IUPAC name | N-(4-aminobutyl)acetamide |
| Molecular formula | C6H14N2O |
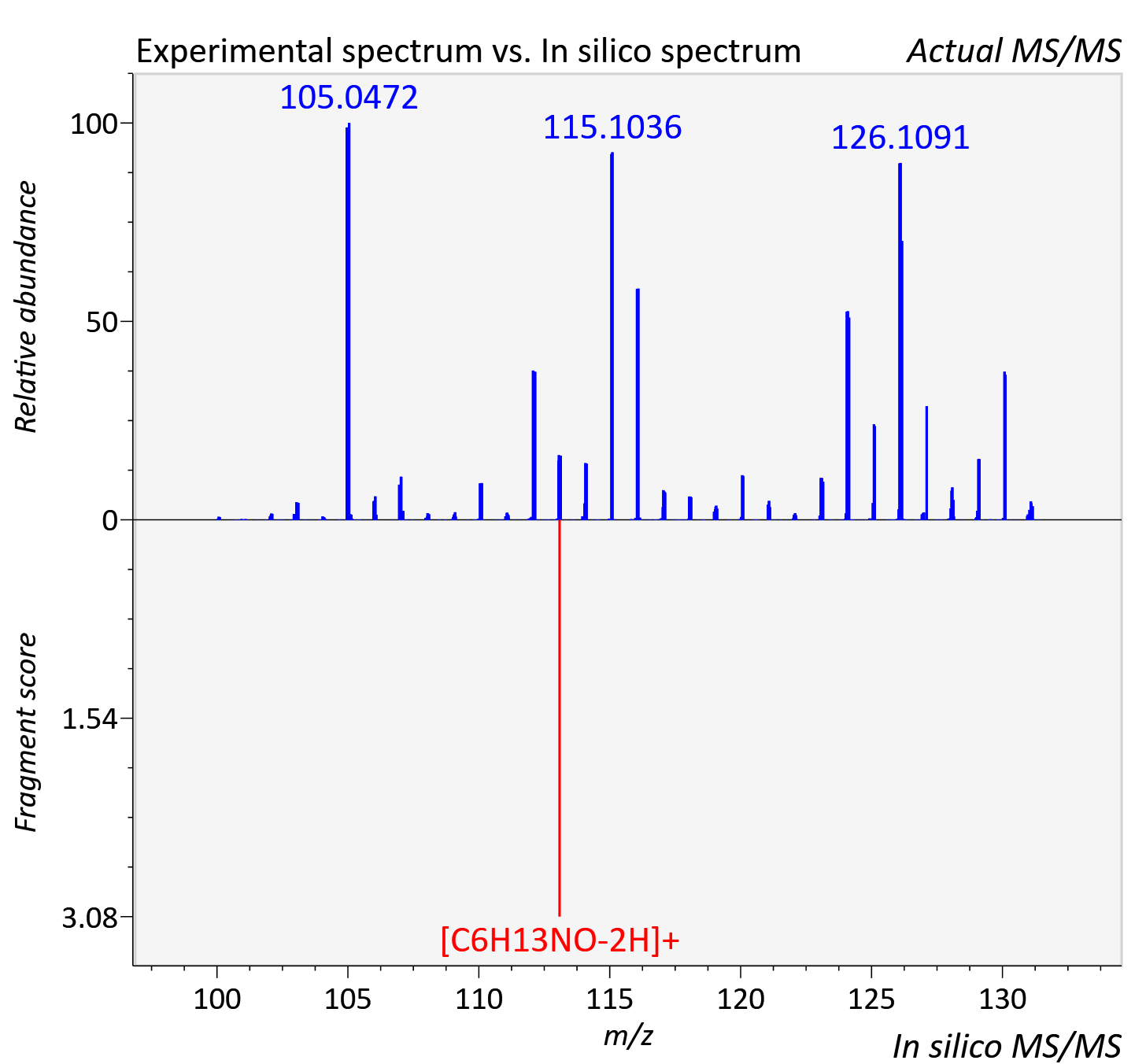
Experimental data
| Retention time | 0.54 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 131.117 | Theoretical mz | 131.118 |
| Error | 6.68 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.4068 |
Identifiers and class information
| Inchi key | KLZGKIDSEJWEDW-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C(NCCCCN)C |
| Superclass | Organic acids and derivatives |
| Class | Carboximidic acids and derivatives |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P24046 | GABRR1 | GABA receptor rho-1 subunit | T99665 | SEA |
| O75899 | GABBR2 | GABA-B receptor | T42446 | SEA |
| P48066 | SLC6A11 | GABA transporter 3 | T38200 | SEA |
| O75899 | GABBR2 | GABA-B receptor | T42446 | SEA |
| Q9NTG7 | SIRT3 | NAD-dependent deacetylase sirtuin 3 | T91959 | SEA |
| Q8IXJ6 | SIRT2 | NAD-dependent deacetylase sirtuin 2 | T83904 | SEA |
| P16278 | GLB1 | Beta-galactosidase (by homology) | T73134 | SEA |
| Q05586 | GRIN1 | Glutamate NMDA receptor; GRIN1/GRIN2B | T62276 | SEA |
| Q12879 | GRIN2A | Glutamate NMDA receptor; GRIN1/GRIN2A | T22285 | SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SEA |
| Q9H4A4 | RNPEP | Aminopeptidase B | T57818 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| Q6QHF9 | PAOX | Polyamine oxidase | T88699 | SEA |
| P48039 | MTNR1A | Melatonin receptor 1A | T97613 | SEA |
| P49286 | MTNR1B | Melatonin receptor 1B | T48268 | SEA |
| P61964 | WDR5 | WD repeat-containing protein 5 | T77393 | SEA |
| Q13224 | GRIN2B | Glutamate [NMDA] receptor subunit epsilon 2 | T76414 | SEA |
| Q04756 | HGFAC | Hepatocyte growth factor activator | T38584 | SEA |
| P09958 | FUR | HUMAN dibasic-processing enzyme | T98945 | SEA |
| Q9Y4G9 | PCSK6 | Proprotein convertase subtilisin/kexin type 6 | T86503 | SEA |
| Q9UN88 | GABRQ | GABA(A) receptor theta | T15161 | SEA |
| Q16348 | PEPT2 | Solute carrier family 15 member 2 | T52067 | SEA |
| Q9NXA8 | SIRT5 | NAD-dependent deacetylase sirtuin-5 | T91940 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T42446 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O75899 | GABBR2 |
| T42446 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | O75899 | GABBR2 |
| T42446 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | O75899 | GABBR2 |
| T42446 | DI0290 | Narcolepsy | [ICD-11: 7A20] | O75899 | GABBR2 |
| T38200 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P48066 | SLC6A11 |
| T42446 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | O75899 | GABBR2 |
| T42446 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | O75899 | GABBR2 |
| T42446 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | O75899 | GABBR2 |
| T42446 | DI0290 | Narcolepsy | [ICD-11: 7A20] | O75899 | GABBR2 |
| T83904 | DI0365 | Retinopathy | [ICD-11: 9B71] | Q8IXJ6 | SIRT2 |
| T73134 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | P16278 | GLB1 |
| T62276 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q05586 | GRIN1 |
| T62276 | DI0182 | HIV-infected patients with tuberculosis | [ICD-11: 1B10-1B14] | Q05586 | GRIN1 |
| T22285 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | Q12879 | GRIN2A |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T33584 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42261 | GRIA1 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T97613 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P48039 | MTNR1A |
| T48268 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P49286 | MTNR1B |
| T76414 | DI0024 | Alpha-1-antitrypsin deficiency | [ICD-11: 5C5A] | Q13224 | GRIN2B |
| T98945 | DI0263 | Middle East Respiratory Syndrome | [ICD-11: 1D64] | P09958 | FUR |
