Compound details
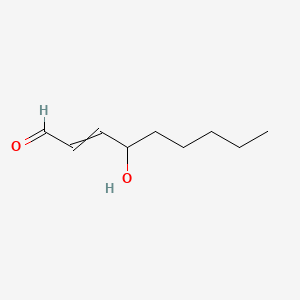
4-Hydroxynonenal
| Compound ID | CDAMM02078 |
|---|---|
| Common name | 4-Hydroxynonenal | IUPAC name | 4-hydroxynon-2-enal |
| Molecular formula | C9H16O2 |
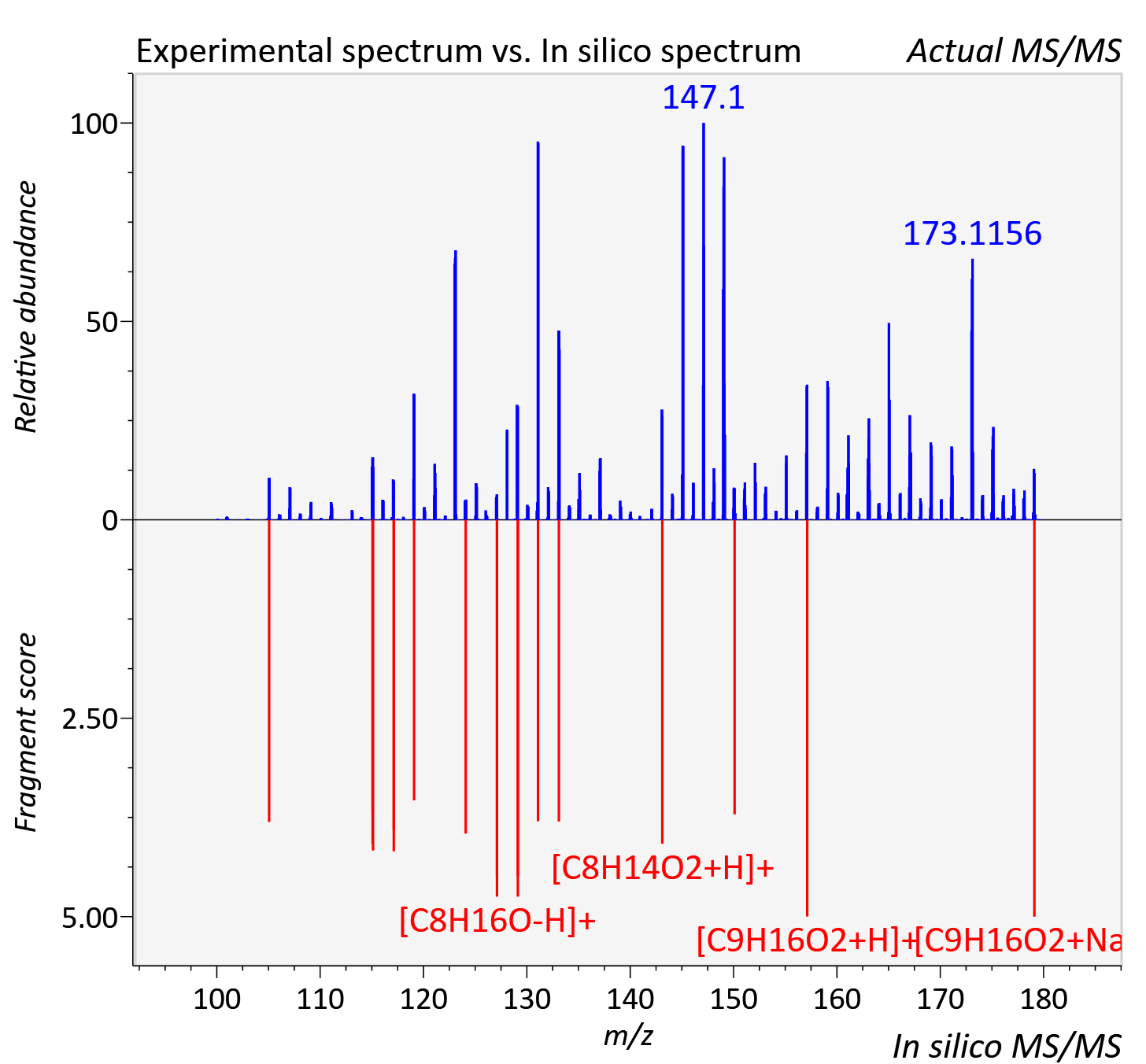
Experimental data
| Retention time | 7.49 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 179.104 | Theoretical mz | 179.104 |
| Error | 0.66 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.3786 |
Identifiers and class information
| Inchi key | JVJFIQYAHPMBBX-FNORWQNLNA-N |
|---|---|
| Smiles | O=CC=CC(O)CCCCC |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| P43116 | PTGER2 | Prostanoid EP2 receptor | T38529 | SEA |
| P35408 | PTGER4 | Prostanoid EP4 receptor | T18876 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| P43088 | PTGFR | Prostanoid FP receptor | T75797 | SEA |
| P43119 | PTGIR | Prostanoid IP receptor | T99954 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P40394 | ADH7 | Alcohol dehydrogenase class IV | T09770 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| Q92959 | SLCO2A1 | Prostaglandin transporter | T15518 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T38529 | DI0003 | Abortion | [ICD-11: JA00] | P43116 | PTGER2 |
| T38529 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43116 | PTGER2 |
| T38529 | DI0121 | Diabetic foot ulcer | [ICD-11: BD54] | P43116 | PTGER2 |
| T38529 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43116 | PTGER2 |
| T38529 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43116 | PTGER2 |
| T38529 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | P43116 | PTGER2 |
| T38529 | DI0413 | Transplant rejection | [ICD-11: NE84] | P43116 | PTGER2 |
| T18876 | DI0037 | Asthma | [ICD-11: CA23] | P35408 | PTGER4 |
| T18876 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P35408 | PTGER4 |
| T18876 | DI0264 | Migraine | [ICD-11: 8A80] | P35408 | PTGER4 |
| T18876 | DI0303 | Non-small-cell lung cancer | [ICD-11: 2C25] | P35408 | PTGER4 |
| T18876 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35408 | PTGER4 |
| T18876 | DI0360 | Rectum cancer | [ICD-11: 2B92] | P35408 | PTGER4 |
| T18876 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35408 | PTGER4 |
| T18876 | DI0414 | Transplanted organ/tissue | [ICD-11: QB63] | P35408 | PTGER4 |
| T75797 | DI0003 | Abortion | [ICD-11: JA00] | P43088 | PTGFR |
| T75797 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43088 | PTGFR |
| T75797 | DI0166 | Glaucoma | [ICD-11: 9C61] | P43088 | PTGFR |
| T99954 | DI0003 | Abortion | [ICD-11: JA00] | P43119 | PTGIR |
| T99954 | DI0102 | Coronary atherosclerosis | [ICD-11: BA52] | P43119 | PTGIR |
| T99954 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P43119 | PTGIR |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
