Compound details
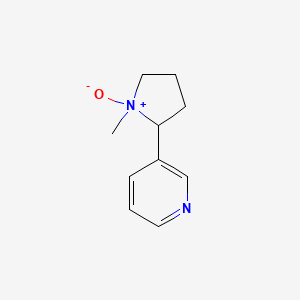
Nicotine-1\'-N-oxide
| Compound ID | CDAMM02066 |
|---|---|
| Common name | Nicotine-1\'-N-oxide | IUPAC name | 3-(1-methyl-1-oxidopyrrolidin-1-ium-2-yl)pyridine |
| Molecular formula | C10H14N2O |
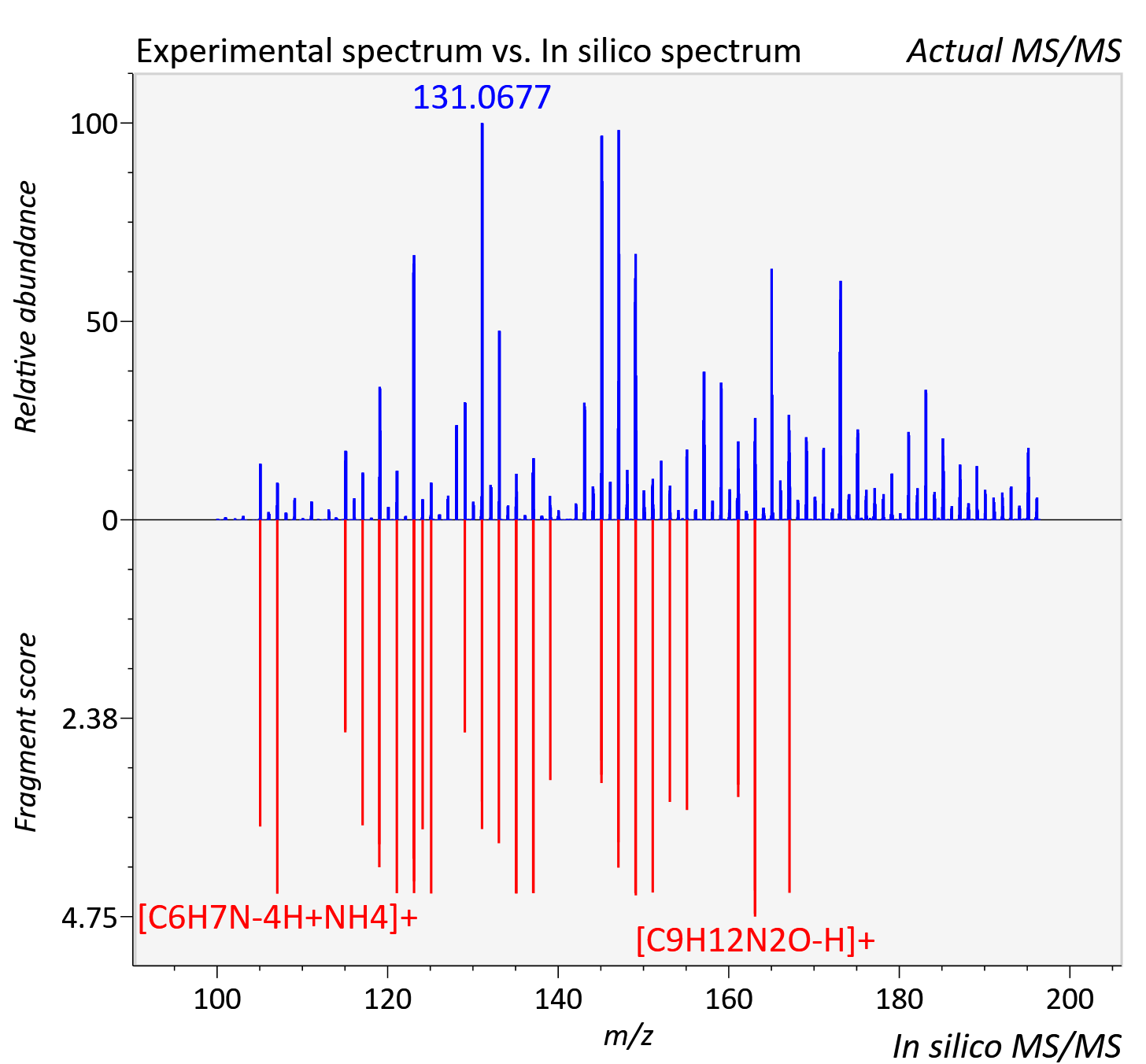
Experimental data
| Retention time | 7.5 |
|---|---|
| Adduct | [M+NH4]+ |
| Actual mz | 196.144 | Theoretical mz | 196.145 |
| Error | 3.69 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.9849 |
Identifiers and class information
| Inchi key | RWFBQHICRCUQJJ-UHFFFAOYNA-N |
|---|---|
| Smiles | [O-][N+]1(C)CCCC1C=2C=NC=CC2 |
| Superclass | Organoheterocyclic compounds |
| Class | Pyridines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43681 | CHRNA4 | Neuronal acetylcholine receptor; alpha4/beta2 | T70967 | SwissTargetPrediction and SEA |
| P36544 | CHRNA7 | Neuronal acetylcholine receptor protein alpha-7 subunit | T34429 | SwissTargetPrediction and SEA |
| P32297 | CHRNA3 | Neuronal acetylcholine receptor; alpha3/beta4 | T74166 | SwissTargetPrediction and SEA |
| P11509 | CYP2A6 | Cytochrome P450 2A6 | T06455 | SEA |
| P55786 | NPEPPS | Puromycin-sensitive aminopeptidase | T15610 | SEA |
| P30926 | CHRNB4 | Neuronal acetylcholine receptor; alpha2/beta4 | T73724 | SwissTargetPrediction and SEA |
| Q15822 | CHRNA2 | Neuronal acetylcholine receptor; alpha2/beta2 | T55815 | SwissTargetPrediction |
| P17787 | CHRNB2 | Nicotinic acetylcholine receptor alpha2/beta2 | T82543 | SEA |
| P02708 | CHRNA1 | Neuronal acetylcholine receptor alpha-1 | T04689 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70967 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P43681 | CHRNA4 |
| T70967 | DI0191 | Hypertensive crisis | [ICD-11: BA03] | P43681 | CHRNA4 |
| T70967 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | P43681 | CHRNA4 |
| T34429 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P36544 | CHRNA7 |
| T34429 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P36544 | CHRNA7 |
| T74166 | DI0105 | Cough | [ICD-11: MD12] | P32297 | CHRNA3 |
| T06455 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P11509 | CYP2A6 |
| T06455 | DI0351 | Psoriasis | [ICD-11: EA90] | P11509 | CYP2A6 |
| T15610 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P55786 | NPEPPS |
| T15610 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | P55786 | NPEPPS |
| T73724 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P30926 | CHRNB4 |
| T55815 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | Q15822 | CHRNA2 |
| T55815 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | Q15822 | CHRNA2 |
| T55815 | DI0166 | Glaucoma | [ICD-11: 9C61] | Q15822 | CHRNA2 |
| T55815 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | Q15822 | CHRNA2 |
| T55815 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | Q15822 | CHRNA2 |
| T82543 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P17787 | CHRNB2 |
| T04689 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P02708 | CHRNA1 |
