Compound details
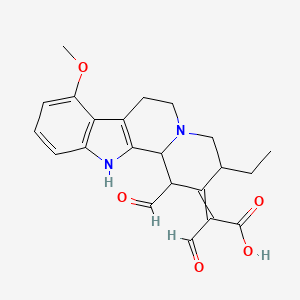
Mitragynalinic acid
| Compound ID | CDAMM02054 |
|---|---|
| Common name | Mitragynalinic acid | IUPAC name | 2-(3-ethyl-1-formyl-8-methoxy-3,4,6,7,12,12b-hexahydro-1H-indolo[2,3-a]quinolizin-2-ylidene)-3-oxopropanoic acid |
| Molecular formula | C22H24N2O5 |
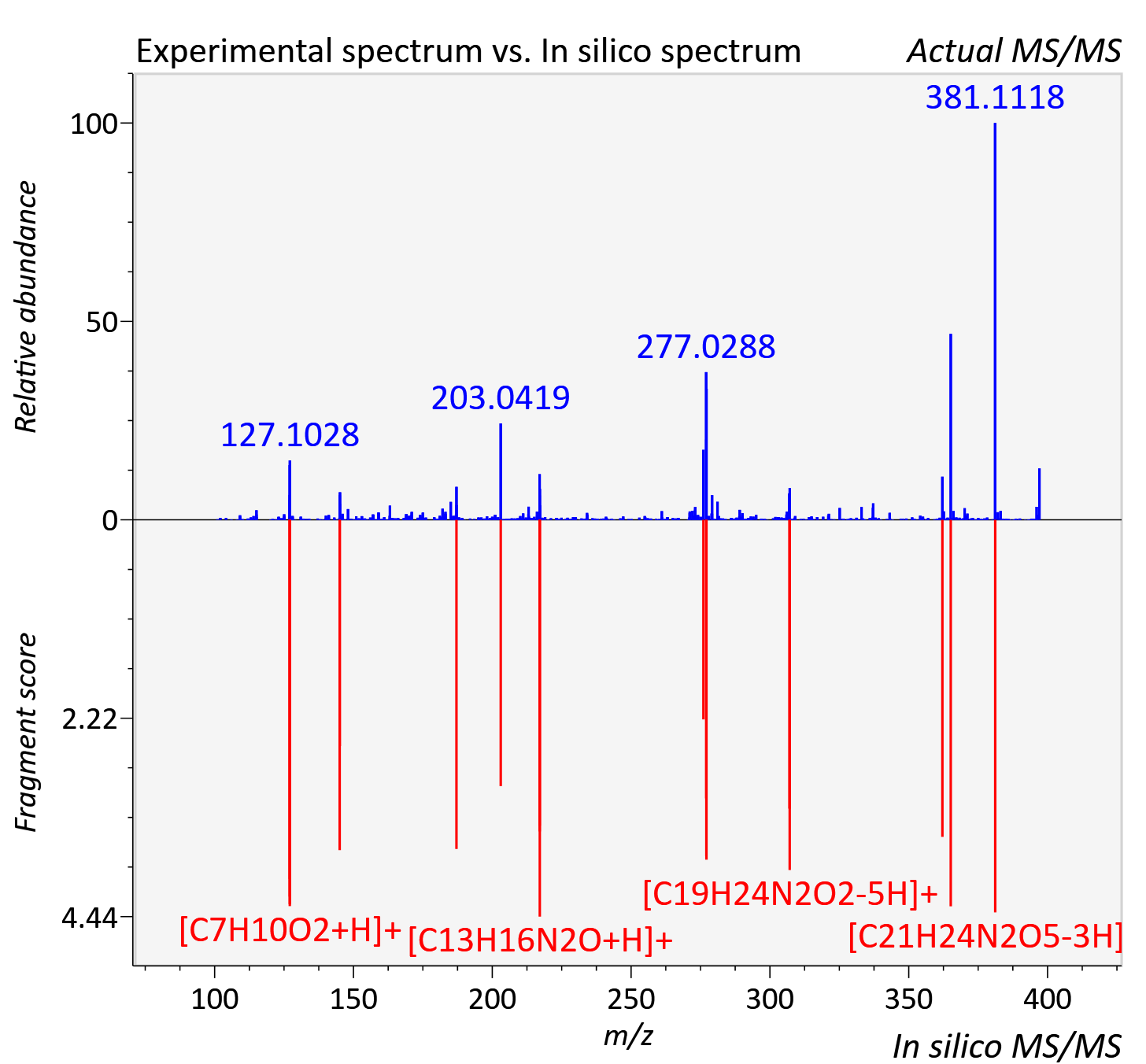
Experimental data
| Retention time | 0.67 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 397.178 | Theoretical mz | 397.176 |
| Error | 3.59 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.5241 |
Identifiers and class information
| Inchi key | BZCUJLGQIQXOMA-SDXDJHTJNA-N |
|---|---|
| Smiles | O=CC(C(=O)O)=C1C(C=O)C2C=3NC=4C=CC=C(OC)C4C3CCN2CC1CC |
| Superclass | Alkaloids and derivatives |
| Class | Corynanthean-type alkaloids |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T20709 | DI0037 | Asthma | [ICD-11: CA23] | P08173 | CHRM4 |
| T20709 | DI0141 | Eye anterior segment structural developmental anomaly | [ICD-11: LA11] | P08173 | CHRM4 |
| T20709 | DI0166 | Glaucoma | [ICD-11: 9C61] | P08173 | CHRM4 |
| T20709 | DI0293 | Nausea/vomiting | [ICD-11: MD90] | P08173 | CHRM4 |
| T20709 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08173 | CHRM4 |
| T79961 | DI0002 | Abnormal micturition | [ICD-11: MF50] | P08912 | CHRM5 |
| T79961 | DI0032 | Anterior uveitis | [ICD-11: 9A96] | P08912 | CHRM5 |
| T79961 | DI0034 | Appearance/behaviour symptom | [ICD-11: MB23] | P08912 | CHRM5 |
| T79961 | DI0037 | Asthma | [ICD-11: CA23] | P08912 | CHRM5 |
| T79961 | DI0061 | Brain disease | [ICD-11: 8C70-8E61] | P08912 | CHRM5 |
| T79961 | DI0071 | Central and peripheral nervous disease | [ICD-11: 8A04-8E7Z] | P08912 | CHRM5 |
| T79961 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P08912 | CHRM5 |
| T79961 | DI0093 | Cognition symptoms/signs/clinical symptom | [ICD-11: MB21] | P08912 | CHRM5 |
| T79961 | DI0105 | Cough | [ICD-11: MD12] | P08912 | CHRM5 |
| T79961 | DI0124 | Digestive system disease | [ICD-11: DE2Z] | P08912 | CHRM5 |
| T79961 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P08912 | CHRM5 |
| T79961 | DI0154 | Functional bladder disorder | [ICD-11: GC50] | P08912 | CHRM5 |
| T79961 | DI0158 | Gastritis | [ICD-11: DA42] | P08912 | CHRM5 |
| T79961 | DI0162 | General examination | [ICD-11: QA00] | P08912 | CHRM5 |
| T79961 | DI0166 | Glaucoma | [ICD-11: 9C61] | P08912 | CHRM5 |
| T79961 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P08912 | CHRM5 |
| T79961 | DI0208 | Infectious gastroenteritis/colitis | [ICD-11: 1A40] | P08912 | CHRM5 |
| T79961 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P08912 | CHRM5 |
| T79961 | DI0307 | Nystagmus | [ICD-11: 9C84] | P08912 | CHRM5 |
| T79961 | DI0315 | Oesophagus motility disorder | [ICD-11: DA21] | P08912 | CHRM5 |
| T79961 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P08912 | CHRM5 |
| T79961 | DI0329 | Pancreatitis | [ICD-11: DC31-DC34] | P08912 | CHRM5 |
| T79961 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P08912 | CHRM5 |
| T79961 | DI0333 | Peptic ulcer | [ICD-11: DA61] | P08912 | CHRM5 |
| T79961 | DI0338 | Polyuria | [ICD-11: MF55] | P08912 | CHRM5 |
| T79961 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P08912 | CHRM5 |
| T79961 | DI0421 | Unspecific substance harmful effect | [ICD-11: NE6Z] | P08912 | CHRM5 |
| T79961 | DI0424 | Urgency | [ICD-11: N.A.] | P08912 | CHRM5 |
| T46185 | DI0037 | Asthma | [ICD-11: CA23] | P08172 | CHRM2 |
| T46185 | DI0166 | Glaucoma | [ICD-11: 9C61] | P08172 | CHRM2 |
| T46185 | DI0278 | Muscle disorder | [ICD-11: FB32-FB3Z] | P08172 | CHRM2 |
| T46185 | DI0333 | Peptic ulcer | [ICD-11: DA61] | P08172 | CHRM2 |
| T46185 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08172 | CHRM2 |
| T46185 | DI0371 | Sebaceous gland disorder | [ICD-11: ED91] | P08172 | CHRM2 |
