Compound details
Sapiol
| Compound ID | CDAMM01970 |
|---|---|
| Common name | Sapiol | IUPAC name | tetratriacontan-1-ol |
| Molecular formula | C34H70O |
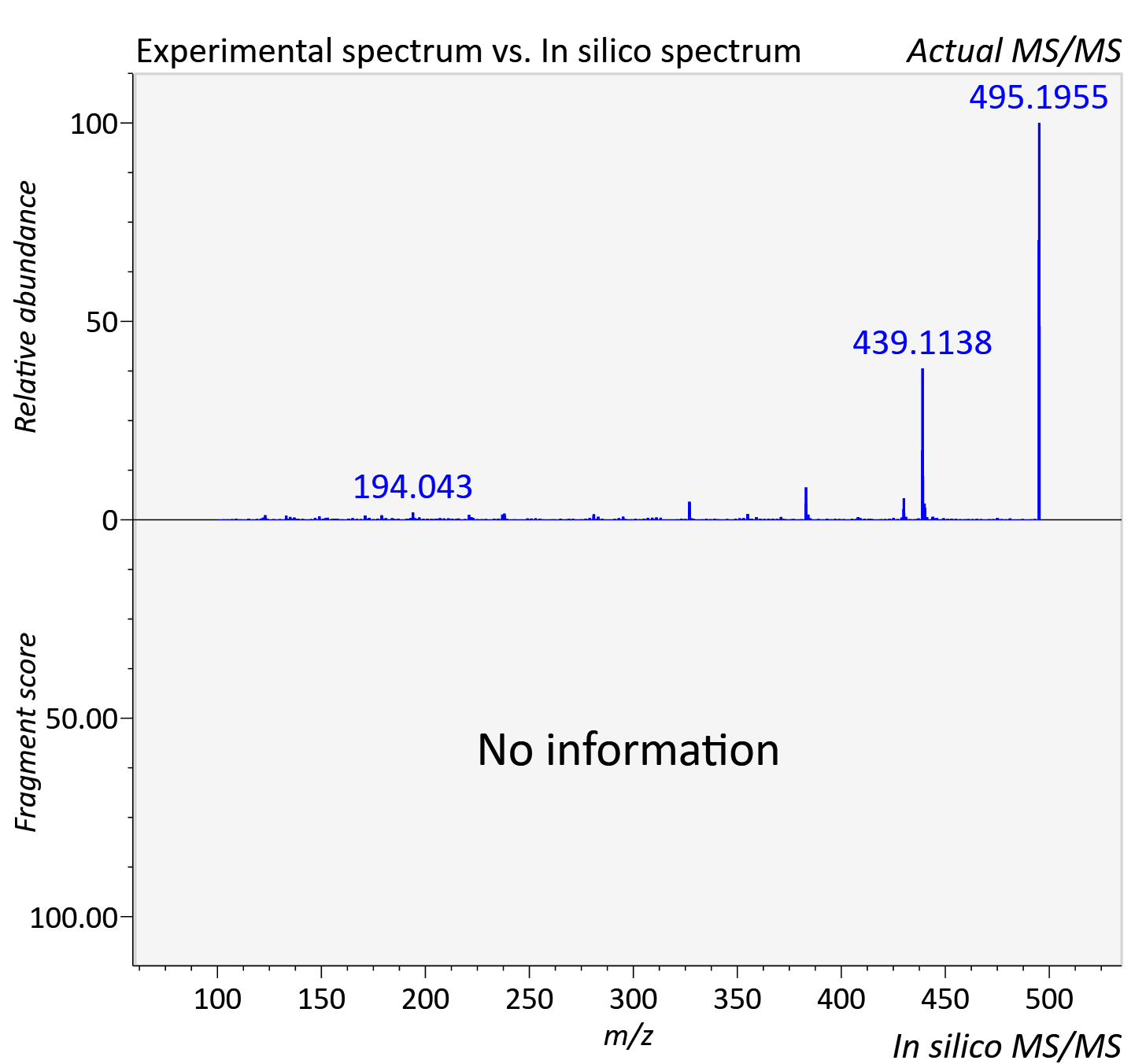
Experimental data
| Retention time | 19.53 |
|---|---|
| Adduct | [2M+H]2+ |
| Actual mz | 495.051 | Theoretical mz | 495.047 |
| Error | 8.81 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.7088 |
Identifiers and class information
| Inchi key | OULAJFUGPPVRBK-UHFFFAOYSA-N |
|---|---|
| Smiles | OCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SEA |
| P04054 | PLA2G1B | Phospholipase A2 group 1B | T31479 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| P10826 | RARB | Retinoic acid receptor beta | T61657 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| O95977 | S1PR4 | Sphingosine 1-phosphate receptor Edg-6 | T17523 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P40394 | ADH7 | Alcohol dehydrogenase class IV | T09770 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| Q9UMJ8 | GBA1 | Lysosomal acid glucosylceramidase | T63243 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T31479 | DI0230 | Leishmaniasis | [ICD-11: 1F54] | P04054 | PLA2G1B |
| T31479 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P04054 | PLA2G1B |
| T31479 | DI0367 | Rosacea | [ICD-11: ED90] | P04054 | PLA2G1B |
| T31479 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P04054 | PLA2G1B |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
| T61657 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P10826 | RARB |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T63243 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | Q9UMJ8 | GBA1 |
