Compound details
Butylcyclohexane
| Compound ID | CDAMM01964 |
|---|---|
| Common name | Butylcyclohexane | IUPAC name | butylcyclohexane |
| Molecular formula | C10H20 |
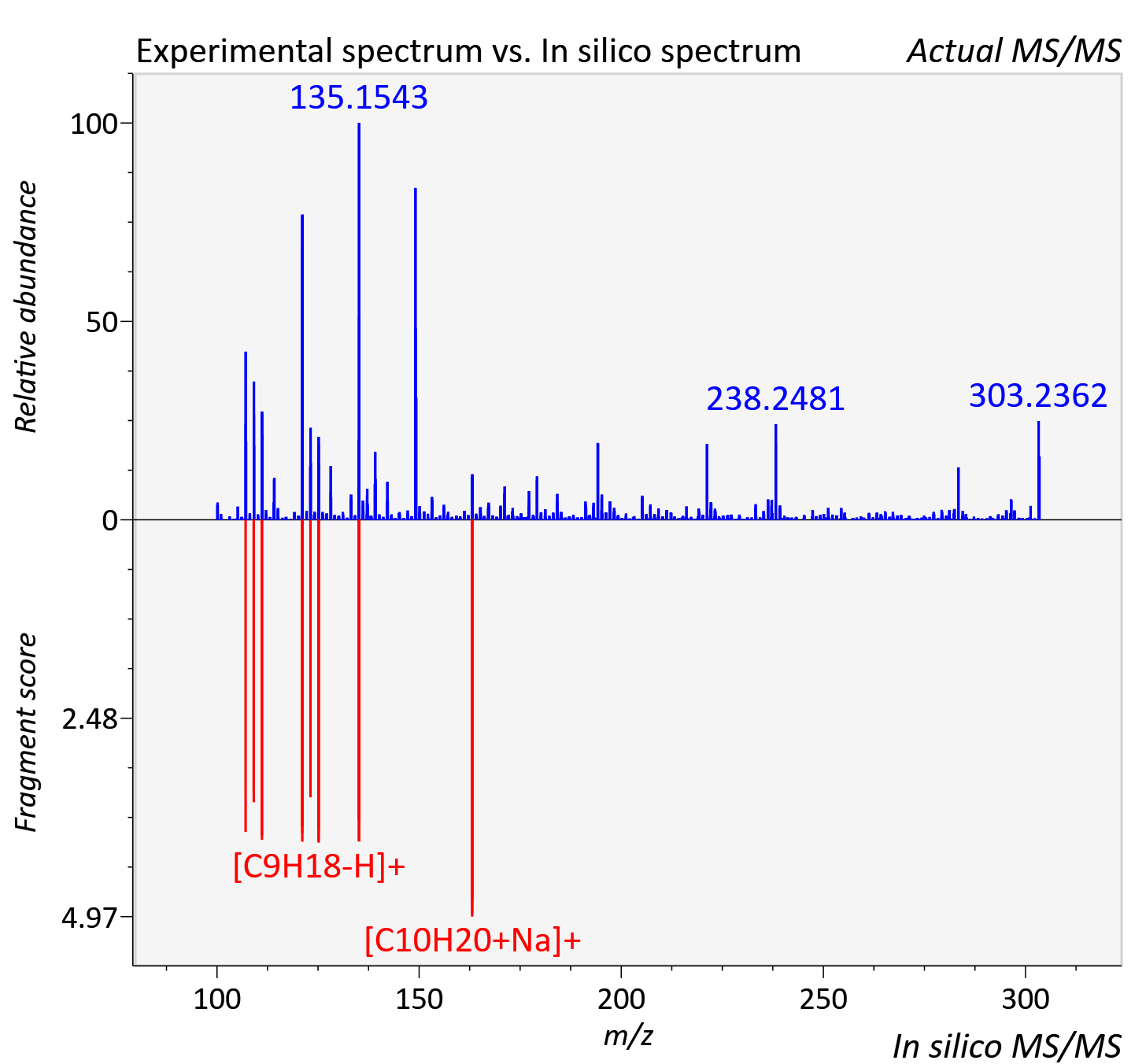
Experimental data
| Retention time | 14.92 |
|---|---|
| Adduct | [2M+Na]+ |
| Actual mz | 303.306 | Theoretical mz | 303.303 |
| Error | 7.85 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.1259 |
Identifiers and class information
| Inchi key | GGBJHURWWWLEQH-UHFFFAOYSA-N |
|---|---|
| Smiles | CCCCC1CCCCC1 |
| Superclass | Hydrocarbons |
| Class | Saturated hydrocarbons |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T61339 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P06280 | GLA |
| T39677 | DI0325 | Pancreas disease | [ICD-11: DC35-DC3Z] | P19835 | CEL |
