Compound details
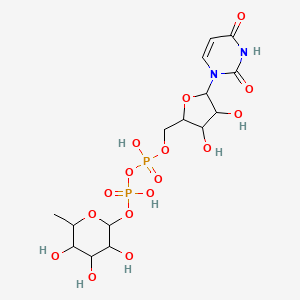
UDP-L-rhamnose
| Compound ID | CDAMM01883 |
|---|---|
| Common name | UDP-L-rhamnose | IUPAC name | [[5-(2,4-dioxopyrimidin-1-yl)-3,4-dihydroxyoxolan-2-yl]methoxy-hydroxyphosphoryl] (3,4,5-trihydroxy-6-methyloxan-2-yl) hydrogen phosphate |
| Molecular formula | C15H24N2O16P2 |
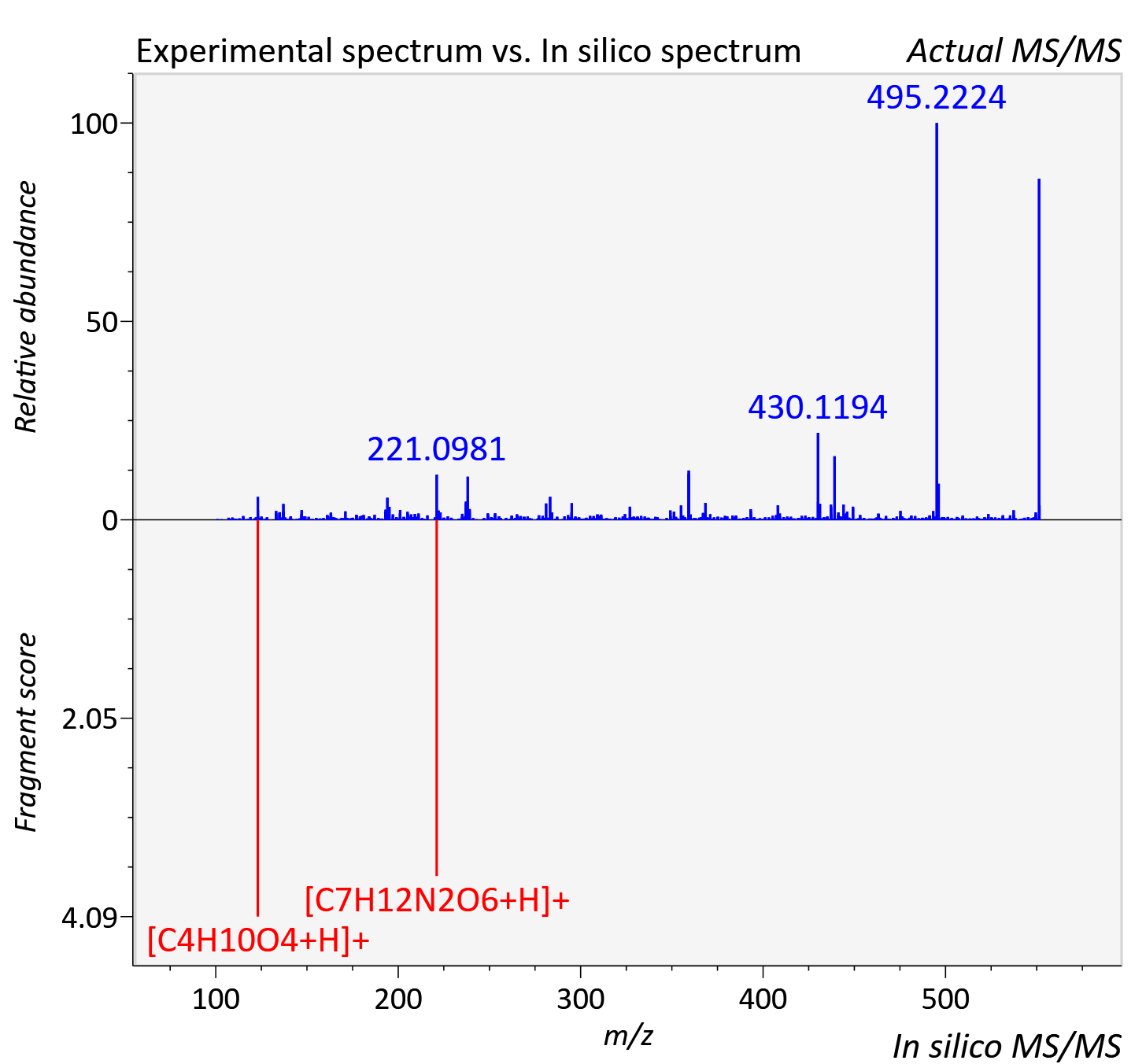
Experimental data
| Retention time | 19.65 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 551.072 | Theoretical mz | 551.067 |
| Error | 9.3 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.2264 |
Identifiers and class information
| Inchi key | DRDCJEIZVLVWNC-SLBWPEPYSA-N |
|---|---|
| Smiles | O=C1C=CN(C(=O)N1)C2OC(COP(=O)(O)OP(=O)(O)OC3OC(C)C(O)C(O)C3O)C(O)C2O |
| Superclass | Nucleosides, nucleotides, and analogues |
| Class | Pyrimidine nucleotides |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| P04818 | TYMS | Thymidylate synthase (by homology) | T98397 | SwissTargetPrediction and SEA |
| P04183 | TK1 | Thymidine kinase, cytosolic | T30081 | SwissTargetPrediction and SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SwissTargetPrediction and SEA |
| P32320 | CDA | Cytidine deaminase (by homology) | T79027 | SEA |
| P55263 | ADK | Adenosine kinase | T91661 | SEA |
| P11142 | HSPA8 | Heat shock cognate 71 kDa protein | T12817 | SwissTargetPrediction |
| P11021 | HSPA5 | 78 kDa glucose-regulated protein | T77594 | SwissTargetPrediction |
| P20839 | IMPDH1 | Inosine-5'-monophosphate dehydrogenase 1 | T40111 | SEA |
| P12268 | IMPDH2 | Inosine-5'-monophosphate dehydrogenase 2 | T89360 | SEA |
| Q9H244 | P2RY12 | Purinergic receptor P2Y12 (by homology) | T46937 | SEA |
| P19971 | TYMP | Thymidine phosphorylase | T59929 | SEA |
| Q9GZQ4 | NMUR2 | Neuromedin-U receptor 2 | T04210 | SEA |
| P47900 | P2RY1 | Purinergic receptor P2Y1 | T67818 | SEA |
| P41231 | P2RY2 | Purinergic receptor P2Y2 | T93515 | SwissTargetPrediction and SEA |
| Q15077 | P2RY6 | Pyrimidinergic receptor P2Y6 | T90782 | SwissTargetPrediction and SEA |
| P63000 | RAC1 | Ras-related C3 botulinum toxin substrate 1 | T88752 | SwissTargetPrediction and SEA |
| P06730 | EIF4E | Eukaryotic translation initation factor (by homology) | T31186 | SwissTargetPrediction |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| P23919 | DTYMK | Thymidylate kinase | T94400 | SwissTargetPrediction and SEA |
| P11172 | UMPS | Uridine 5'-monophosphate synthase | T15851 | SwissTargetPrediction and SEA |
| O94759 | TRPM2 | Transient receptor potential cation channel subfamily M member 2 | T78205 | SEA |
| Q15391 | P2RY14 | Purinergic receptor P2Y14 | T16555 | SwissTargetPrediction and SEA |
| Q05823 | RNASEL | RNase L | T25200 | SEA |
| P51575 | P2RX1 | P2X purinoceptor 1 | T69091 | SEA |
| P28340 | POLD1 | DNA polymerase delta subunit 1 | T02258 | SwissTargetPrediction |
| P28340 | POLD1 | DNA polymerase delta subunit 1 | T02258 | SEA |
| Q99571 | P2RX4 | P2X purinoceptor 4 | T60330 | SEA |
| P11717 | IGF2R | Insulin-like growth factor II receptor | T59550 | SEA |
| Q13304 | GPR17 | Uracil nucleotide/cysteinyl leukotriene receptor | T33902 | SEA |
| Q9Y4W7 | PARG | Poly(ADP-ribose) glycohydrolase | T66935 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T98397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P04818 | TYMS |
| T98397 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P04818 | TYMS |
| T30081 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P04183 | TK1 |
| T30081 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | P04183 | TK1 |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T79027 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P32320 | CDA |
| T91661 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P55263 | ADK |
| T77594 | DI0252 | Melanoma | [ICD-11: 2C30] | P11021 | HSPA5 |
| T77594 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P11021 | HSPA5 |
| T40111 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P20839 | IMPDH1 |
| T40111 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P20839 | IMPDH1 |
| T40111 | DI0250 | Mature B-cell lymphoma | [ICD-11: 2A85] | P20839 | IMPDH1 |
| T40111 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P20839 | IMPDH1 |
| T40111 | DI0413 | Transplant rejection | [ICD-11: NE84] | P20839 | IMPDH1 |
| T89360 | DI0413 | Transplant rejection | [ICD-11: NE84] | P12268 | IMPDH2 |
| T46937 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | Q9H244 | P2RY12 |
| T46937 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | Q9H244 | P2RY12 |
| T59929 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P19971 | TYMP |
| T93515 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P41231 | P2RY2 |
| T90782 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q15077 | P2RY6 |
| T88752 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P63000 | RAC1 |
| T02258 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P28340 | POLD1 |
| T02258 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P28340 | POLD1 |
| T59550 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | P11717 | IGF2R |
| T59550 | DI0368 | Sarcoidosis | [ICD-11: 4B20] | P11717 | IGF2R |
