Compound details
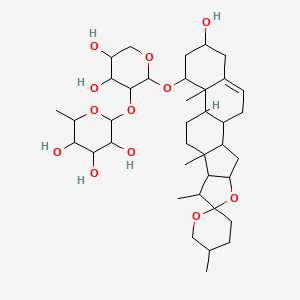
Alliospiroside A
| Compound ID | CDAMM01877 |
|---|---|
| Common name | Alliospiroside A | IUPAC name | 2-[4,5-dihydroxy-2-(16-hydroxy-5',7,9,13-tetramethylspiro[5-oxapentacyclo[10.8.0.02,9.04,8.013,18]icos-18-ene-6,2\'-oxane]-14-yl)oxyoxan-3-yl]oxy-6-methyloxane-3,4,5-triol |
| Molecular formula | C38H60O12 |
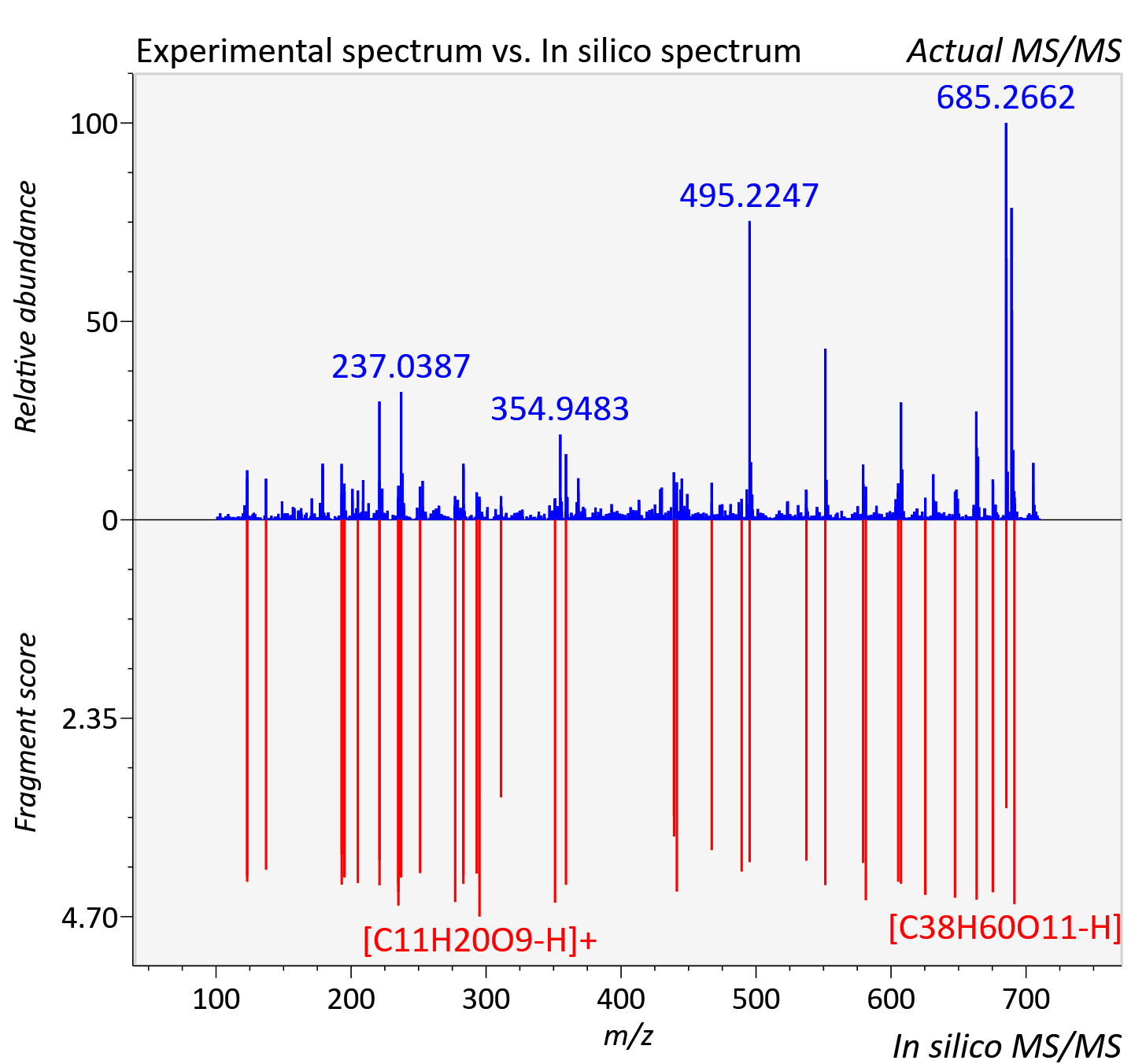
Experimental data
| Retention time | 22.9 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 709.415 | Theoretical mz | 709.415 |
| Error | 0.11 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.5372 |
Identifiers and class information
| Inchi key | SKHJNNFXCKTDBG-UHFFFAOYNA-N |
|---|---|
| Smiles | OC1CC2=CCC3C(CCC4(C)C3CC5OC6(OCC(C)CC6)C(C)C54)C2(C)C(OC7OCC(O)C(O)C7OC8OC(C)C(O)C(O)C8O)C1 |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
External sources
| HMDB ID | |
|---|---|
| KNApSAcK ID | |
| FooDB ID | FDB007854 |
| DrugBank ID | |
| ChEBI ID | |
| Pubchem Compound ID | 3509806 |
| PlantCyc ID | |
| UNPD ID | UNPD165071 |
| Coconut ID | CNP0295047 |
| LipidMAPS ID | |
| NANPDB ID | NANPDB_2198 |
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T25317 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q15125 | EBP |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
