Compound details
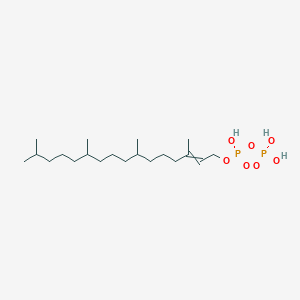
Phytyl diphosphate
| Compound ID | CDAMM01870 |
|---|---|
| Common name | Phytyl diphosphate | IUPAC name | phosphono 3,7,11,15-tetramethylhexadec-2-enyl hydrogen phosphate |
| Molecular formula | C20H42O7P2 |
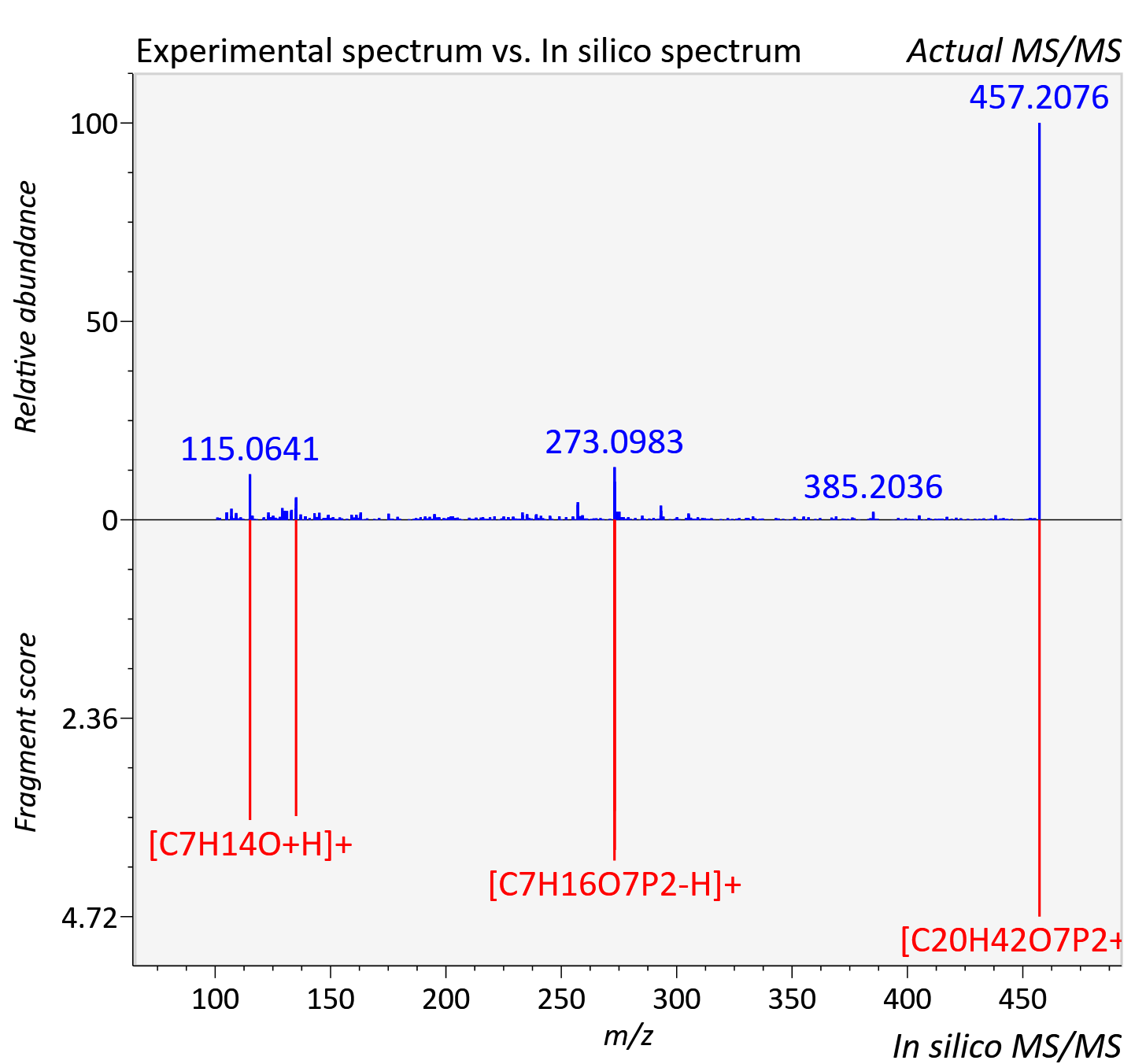
Experimental data
| Retention time | 6.16 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 457.243 | Theoretical mz | 457.248 |
| Error | 11.1 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.5449 |
Identifiers and class information
| Inchi key | ITPLBNCCPZSWEU-HMMYKYKNNA-N |
|---|---|
| Smiles | O=P(O)(O)OP(=O)(O)OCC=C(C)CCCC(C)CCCC(C)CCCC(C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SwissTargetPrediction and SEA |
| P53609 | PGGT1B | Geranylgeranyl transferase type I | T76396 | SwissTargetPrediction |
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P53602 | MVD | Diphosphomevalonate decarboxylase | T96862 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
| T76396 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P53609 | PGGT1B |
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
