Compound details
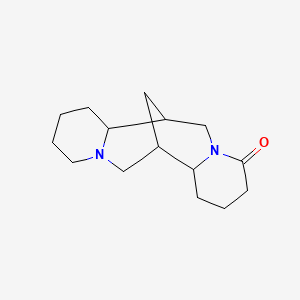
Lupanine
| Compound ID | CDAMM01861 |
|---|---|
| Common name | Lupanine | IUPAC name | 7,15-diazatetracyclo[7.7.1.02,7.010,15]heptadecan-6-one |
| Molecular formula | C15H24N2O |
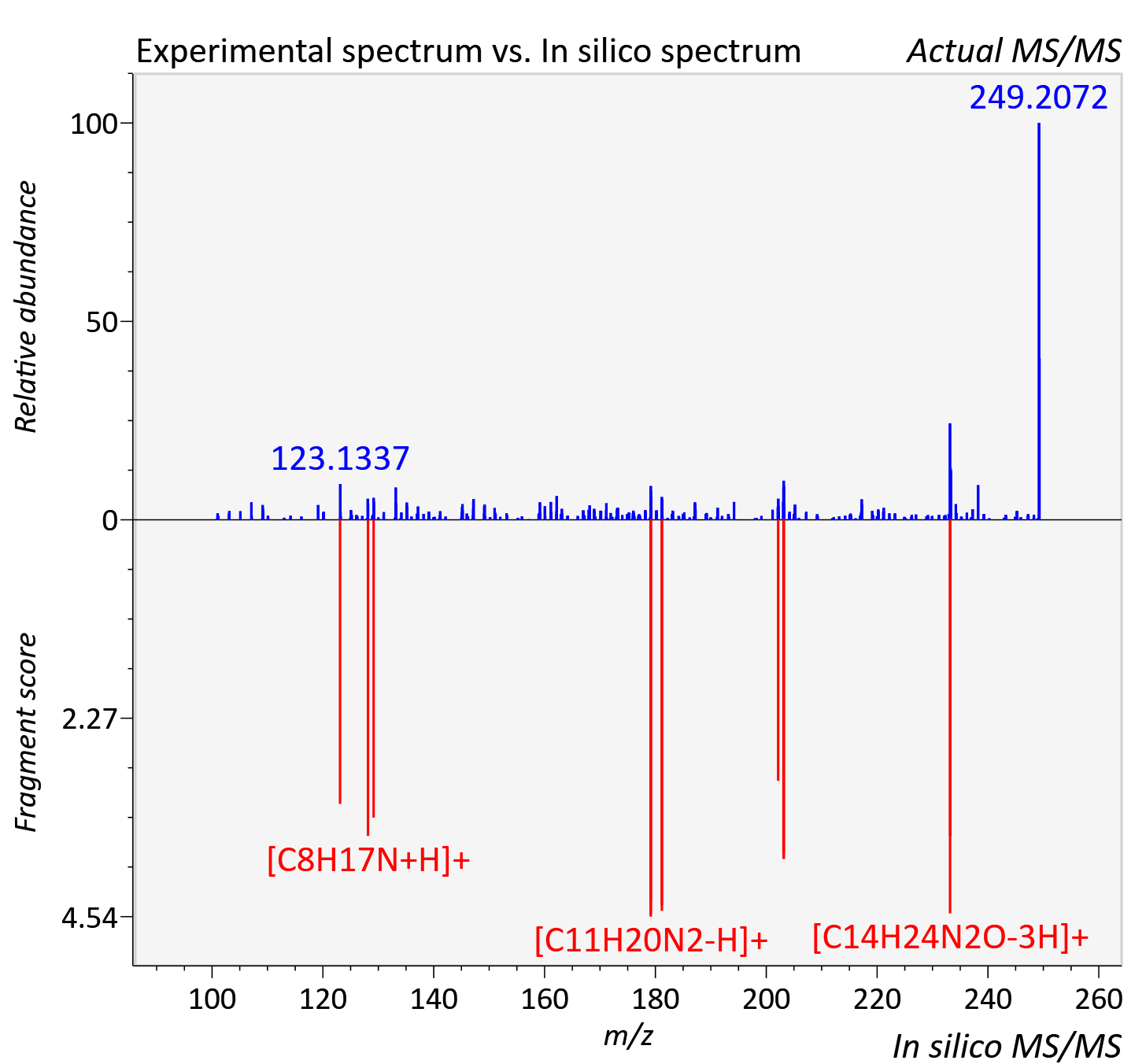
Experimental data
| Retention time | 9.67 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 249.195 | Theoretical mz | 249.196 |
| Error | 6.32 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0623 |
Identifiers and class information
| Inchi key | JYIJIIVLEOETIQ-WDOYOSBINA-N |
|---|---|
| Smiles | O=C1N2CC3CC(CN4CCCCC43)C2CCC1 |
| Superclass | Alkaloids and derivatives |
| Class | Lupin alkaloids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43681 | CHRNA4 | Neuronal acetylcholine receptor; alpha4/beta2 | T70967 | SwissTargetPrediction |
| P36544 | CHRNA7 | Neuronal acetylcholine receptor protein alpha-7 subunit | T34429 | SwissTargetPrediction |
| P32297 | CHRNA3 | Neuronal acetylcholine receptor; alpha3/beta4 | T74166 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70967 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P43681 | CHRNA4 |
| T70967 | DI0191 | Hypertensive crisis | [ICD-11: BA03] | P43681 | CHRNA4 |
| T70967 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | P43681 | CHRNA4 |
| T34429 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P36544 | CHRNA7 |
| T34429 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P36544 | CHRNA7 |
| T74166 | DI0105 | Cough | [ICD-11: MD12] | P32297 | CHRNA3 |
