Compound details
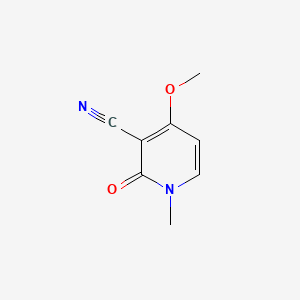
Ricinine
| Compound ID | CDAMM01851 |
|---|---|
| Common name | Ricinine | IUPAC name | 4-methoxy-1-methyl-2-oxopyridine-3-carbonitrile |
| Molecular formula | C8H8N2O2 |
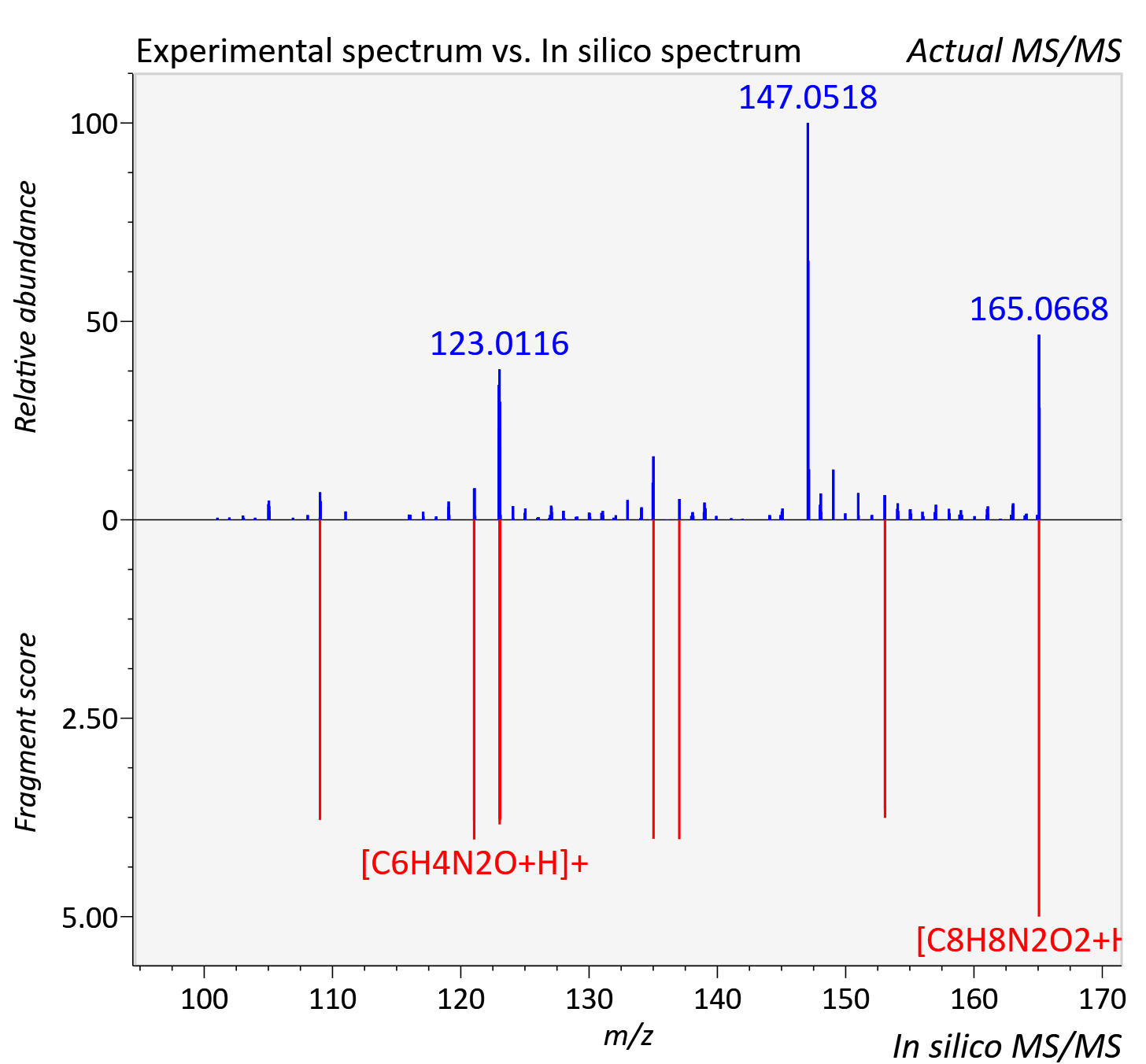
Experimental data
| Retention time | 18.86 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 165.065 | Theoretical mz | 165.066 |
| Error | 5.31 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2138 |
Identifiers and class information
| Inchi key | PETSAYFQSGAEQY-UHFFFAOYSA-N |
|---|---|
| Smiles | N#CC1=C(OC)C=CN(C1=O)C |
| Superclass | Organoheterocyclic compounds |
| Class | Pyridines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P22303 | ACHE | Acetylcholinesterase | T30082 | SwissTargetPrediction |
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SwissTargetPrediction |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SwissTargetPrediction |
| P29475 | NOS1 | Nitric-oxide synthase, brain | T16117 | SwissTargetPrediction |
| P35228 | NOS2 | Nitric oxide synthase, inducible | T02703 | SwissTargetPrediction |
| P29474 | NOS3 | Nitric-oxide synthase, endothelial | T06046 | SwissTargetPrediction |
| P09874 | PARP1 | Poly [ADP-ribose] polymerase-1 | T06273 | SwissTargetPrediction |
| P00915 | CA1 | Carbonic anhydrase I | T13201 | SwissTargetPrediction |
| P00918 | CA2 | Carbonic anhydrase II | T20401 | SwissTargetPrediction |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SwissTargetPrediction |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SwissTargetPrediction |
| P49798 | RGS4 | Regulator of G-protein signaling 4 | T93580 | SwissTargetPrediction |
| P14174 | MIF | Macrophage migration inhibitory factor | T39977 | SwissTargetPrediction |
| P11474 | ESRRA | Estrogen-related receptor alpha | T72841 | SEA |
| Q99685 | MGLL | Monoglyceride lipase | T18664 | SwissTargetPrediction |
| P49841 | GSK3B | Glycogen synthase kinase-3 beta | T70977 | SwissTargetPrediction |
| Q09472 | EP300 | Histone acetyltransferase p300 | T25956 | SwissTargetPrediction |
| Q92831 | KAT2B | Histone acetyltransferase PCAF | T16902 | SwissTargetPrediction |
| O95696 | BRD1 | Bromodomain-containing protein 1 | T35809 | SEA |
| Q9NPI1 | BRD7 | Bromodomain-containing protein 7 | T23984 | SEA |
| Q9H8M2 | BRD9 | Bromodomain-containing protein 9 | T46216 | SEA |
| P04271 | S100B | S100 calcium-binding protein B | T87206 | SEA |
| Q9ULD4 | KIAA1286 | Bromodomain and PHD finger containing 3 | T27466 | SEA |
| O15164 | TRIM24 | Tripartite motif-containing 24 | T56108 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T30082 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P22303 | ACHE |
| T30082 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22303 | ACHE |
| T30082 | DI0282 | Myasthenia gravis | [ICD-11: 8C6Y] | P22303 | ACHE |
| T30082 | DI0313 | Oesophageal/gastroduodenal disorder | [ICD-11: DD90] | P22303 | ACHE |
| T30082 | DI0332 | Pediculosis | [ICD-11: 1G00] | P22303 | ACHE |
| T30082 | DI0421 | Unspecific substance harmful effect | [ICD-11: NE6Z] | P22303 | ACHE |
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T16117 | DI0173 | Headache | [ICD-11: 8A80-8A84] | P29475 | NOS1 |
| T16117 | DI0264 | Migraine | [ICD-11: 8A80] | P29475 | NOS1 |
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T06046 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P29474 | NOS3 |
| T06273 | DI0006 | Acquired cutaneous blood vessel malformation | [ICD-11: EF20] | P09874 | PARP1 |
| T06273 | DI0143 | Fallopian tube cancer | [ICD-11: 2C74] | P09874 | PARP1 |
| T06273 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P09874 | PARP1 |
| T06273 | DI0334 | Peritoneal cancer | [ICD-11: 2C51] | P09874 | PARP1 |
| T13201 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00915 | CA1 |
| T13201 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00915 | CA1 |
| T20401 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P00918 | CA2 |
| T20401 | DI0137 | Essential hypertension | [ICD-11: BA00] | P00918 | CA2 |
| T20401 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00918 | CA2 |
| T20401 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P00918 | CA2 |
| T20401 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P00918 | CA2 |
| T20401 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00918 | CA2 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T39977 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P14174 | MIF |
| T72841 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P11474 | ESRRA |
| T18664 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q99685 | MGLL |
| T18664 | DI0409 | Tic disorder | [ICD-11: 8A05] | Q99685 | MGLL |
| T70977 | DI0288 | Myotonic disorder | [ICD-11: 8C71] | P49841 | GSK3B |
| T25956 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q09472 | EP300 |
| T25956 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q09472 | EP300 |
| T46216 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | Q9H8M2 | BRD9 |
| T87206 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P04271 | S100B |
